Probe CUST_45208_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45208_PI426222305 | JHI_St_60k_v1 | DMT400010866 | GAATCAGTCACACATTTCTCTCTGTCTTCTTCTCTGAAACATTTTTCCTATTTTGAAGGA |
All Microarray Probes Designed to Gene DMG400004253
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45208_PI426222305 | JHI_St_60k_v1 | DMT400010866 | GAATCAGTCACACATTTCTCTCTGTCTTCTTCTCTGAAACATTTTTCCTATTTTGAAGGA |
Microarray Signals from CUST_45208_PI426222305
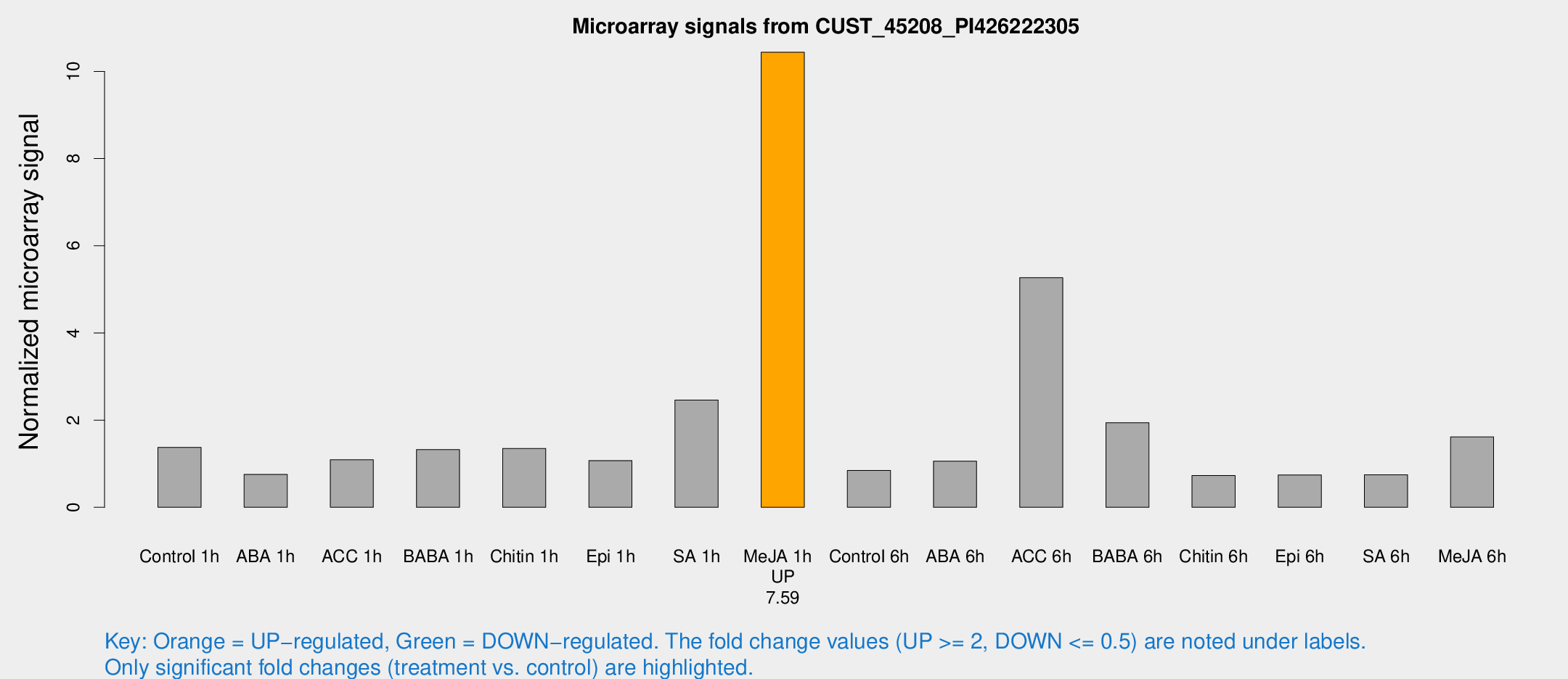
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.6906 | 3.47822 | 1.3758 | 0.5671 |
| ABA 1h | 4.99938 | 2.89756 | 0.753535 | 0.429366 |
| ACC 1h | 9.79551 | 3.81798 | 1.0901 | 0.506085 |
| BABA 1h | 11.2396 | 4.35413 | 1.32286 | 0.515969 |
| Chitin 1h | 10.0975 | 3.0985 | 1.35111 | 0.509736 |
| Epi 1h | 7.60323 | 3.05255 | 1.07207 | 0.494202 |
| SA 1h | 21.3719 | 7.03945 | 2.4618 | 0.79385 |
| Me-JA 1h | 66.6972 | 10.8423 | 10.4397 | 0.793104 |
| Control 6h | 6.50084 | 3.13666 | 0.844995 | 0.414097 |
| ABA 6h | 9.11923 | 3.26941 | 1.05916 | 0.430401 |
| ACC 6h | 50.3524 | 13.8284 | 5.26832 | 1.9133 |
| BABA 6h | 20.5384 | 9.80163 | 1.94119 | 0.959662 |
| Chitin 6h | 5.91235 | 3.42591 | 0.72843 | 0.421891 |
| Epi 6h | 6.4349 | 3.59253 | 0.742175 | 0.411437 |
| SA 6h | 5.63782 | 3.27341 | 0.747749 | 0.434065 |
| Me-JA 6h | 13.8204 | 4.01962 | 1.61517 | 0.712104 |
Source Transcript PGSC0003DMT400010866 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G11820.2 | +3 | 0.0 | 786 | 382/459 (83%) | hydroxymethylglutaryl-CoA synthase / HMG-CoA synthase / 3-hydroxy-3-methylglutaryl coenzyme A synthase | chr4:7109124-7111901 REVERSE LENGTH=461 |
