Probe CUST_4509_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4509_PI426222305 | JHI_St_60k_v1 | DMT400070545 | TCAATGGATTTATACGATACTGTTAATGGGGTTTGGTTAAAAACCCGATCTGTTCCCGGC |
All Microarray Probes Designed to Gene DMG400027417
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4509_PI426222305 | JHI_St_60k_v1 | DMT400070545 | TCAATGGATTTATACGATACTGTTAATGGGGTTTGGTTAAAAACCCGATCTGTTCCCGGC |
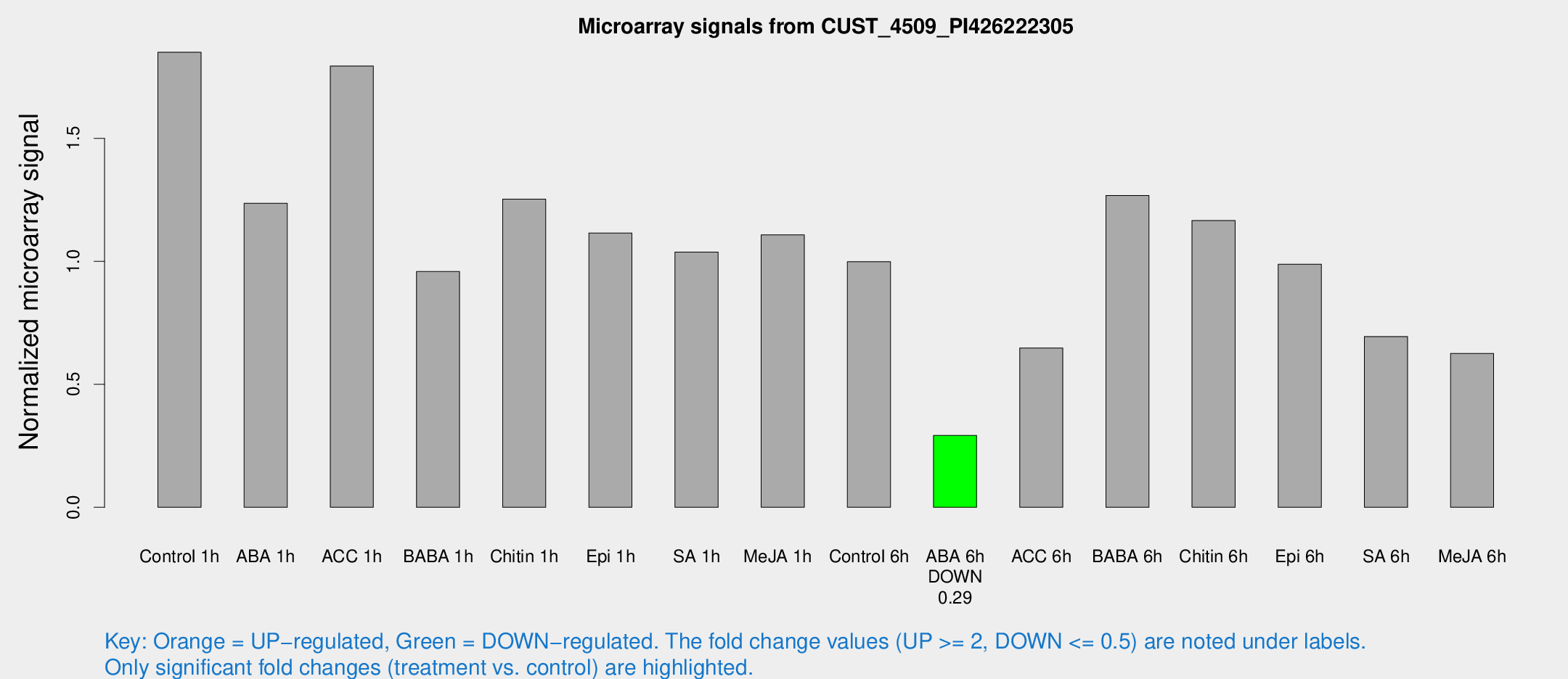
Microarray Signals from CUST_4509_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 150.637 | 22.2326 | 1.85038 | 0.175555 |
| ABA 1h | 88.1293 | 9.94248 | 1.23628 | 0.144523 |
| ACC 1h | 148.6 | 16.9801 | 1.79413 | 0.306532 |
| BABA 1h | 77.7387 | 17.2926 | 0.958827 | 0.142396 |
| Chitin 1h | 93.2786 | 19.6565 | 1.25284 | 0.282258 |
| Epi 1h | 78.5329 | 12.2069 | 1.1154 | 0.139167 |
| SA 1h | 85.2424 | 6.47677 | 1.03742 | 0.146432 |
| Me-JA 1h | 72.4723 | 6.16579 | 1.10801 | 0.188578 |
| Control 6h | 91.6467 | 28.4001 | 0.998757 | 0.337809 |
| ABA 6h | 24.9948 | 3.85551 | 0.292585 | 0.0467053 |
| ACC 6h | 65.7397 | 20.8136 | 0.647953 | 0.141173 |
| BABA 6h | 145.186 | 74.799 | 1.2678 | 0.71636 |
| Chitin 6h | 99.8364 | 10.6762 | 1.16585 | 0.0913009 |
| Epi 6h | 94.0614 | 23.9151 | 0.988691 | 0.264776 |
| SA 6h | 56.7626 | 11.5189 | 0.694497 | 0.1427 |
| Me-JA 6h | 54.4119 | 14.6564 | 0.625837 | 0.15848 |
Source Transcript PGSC0003DMT400070545 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G26960.1 | +1 | 8e-132 | 388 | 224/411 (55%) | Galactose oxidase/kelch repeat superfamily protein | chr5:9484908-9486149 REVERSE LENGTH=413 |
