Probe CUST_45053_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45053_PI426222305 | JHI_St_60k_v1 | DMT400027349 | TATATAAAAGGAGTTCGAATTTAAACCCATGGTTGAATCCACCACGTATGATTGCCATTT |
All Microarray Probes Designed to Gene DMG400010547
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45053_PI426222305 | JHI_St_60k_v1 | DMT400027349 | TATATAAAAGGAGTTCGAATTTAAACCCATGGTTGAATCCACCACGTATGATTGCCATTT |
| CUST_45029_PI426222305 | JHI_St_60k_v1 | DMT400027348 | GTAAAGAAGGTATTTTACCTTTATAACTGGAAGACTGGGCTTGGTCCTATTTATGGGTGA |
Microarray Signals from CUST_45053_PI426222305
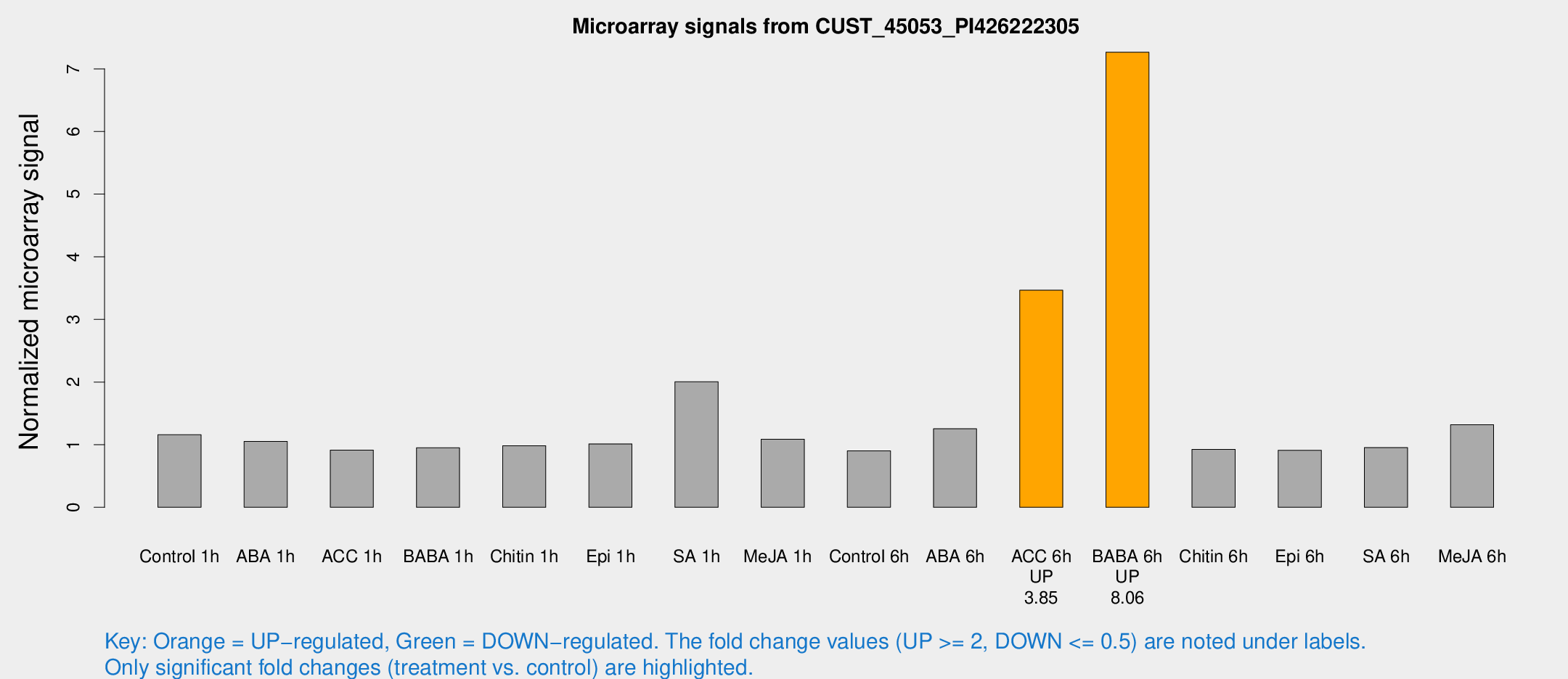
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8.0254 | 3.21011 | 1.15853 | 0.551962 |
| ABA 1h | 5.98724 | 3.10192 | 1.05298 | 0.553472 |
| ACC 1h | 5.98538 | 3.47186 | 0.913698 | 0.529107 |
| BABA 1h | 5.83246 | 3.37921 | 0.950041 | 0.550318 |
| Chitin 1h | 5.63731 | 3.24341 | 0.983815 | 0.562785 |
| Epi 1h | 5.58887 | 3.22496 | 1.01258 | 0.583072 |
| SA 1h | 19.1817 | 11.7381 | 2.00425 | 1.86446 |
| Me-JA 1h | 5.65918 | 3.28262 | 1.08665 | 0.629186 |
| Control 6h | 5.75182 | 3.33777 | 0.901132 | 0.522891 |
| ABA 6h | 9.79425 | 3.89323 | 1.25331 | 0.586195 |
| ACC 6h | 26.2058 | 4.67466 | 3.46551 | 0.835946 |
| BABA 6h | 52.0492 | 4.79692 | 7.26556 | 0.753848 |
| Chitin 6h | 6.27859 | 3.63715 | 0.92591 | 0.536191 |
| Epi 6h | 6.57903 | 3.84345 | 0.909663 | 0.527587 |
| SA 6h | 6.00773 | 3.48431 | 0.95312 | 0.552409 |
| Me-JA 6h | 9.85114 | 4.18702 | 1.31803 | 0.612769 |
Source Transcript PGSC0003DMT400027349 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G16720.1 | +1 | 2e-18 | 83 | 54/134 (40%) | TOXICOS EN LEVADURA 2 | chr3:5692880-5693794 FORWARD LENGTH=304 |
