Probe CUST_44985_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44985_PI426222305 | JHI_St_60k_v1 | DMT400011222 | TAACACAGCAGCTTCTGCAGCATCACTTCTGTACACACGTTAATCTTTGAACAGGTCTGC |
All Microarray Probes Designed to Gene DMG400004395
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44985_PI426222305 | JHI_St_60k_v1 | DMT400011222 | TAACACAGCAGCTTCTGCAGCATCACTTCTGTACACACGTTAATCTTTGAACAGGTCTGC |
Microarray Signals from CUST_44985_PI426222305
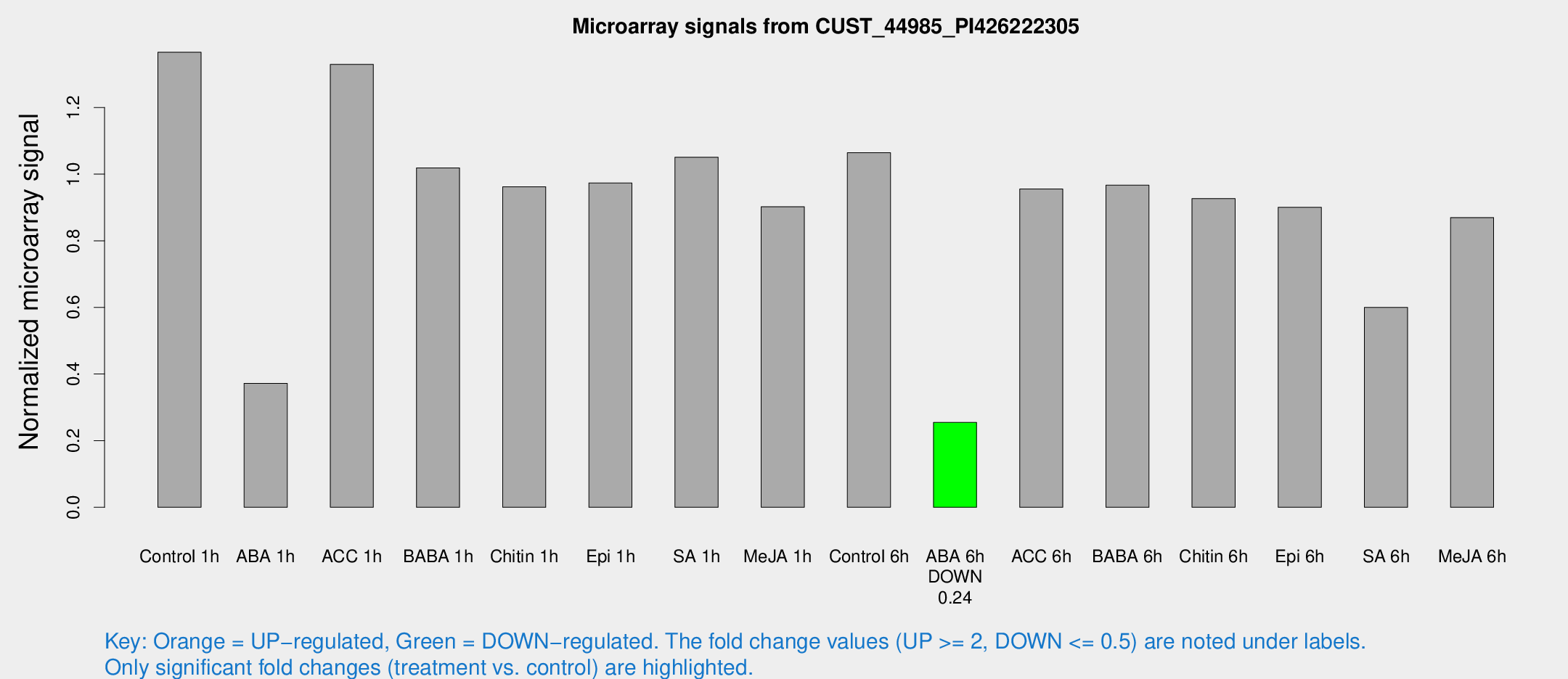
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 141.09 | 14.1862 | 1.36585 | 0.179118 |
| ABA 1h | 38.4414 | 14.6478 | 0.372176 | 0.179264 |
| ACC 1h | 139.966 | 9.26423 | 1.32946 | 0.0884345 |
| BABA 1h | 102.653 | 15.2596 | 1.01876 | 0.0731818 |
| Chitin 1h | 91.5896 | 16.8734 | 0.96224 | 0.244983 |
| Epi 1h | 88.1706 | 13.1412 | 0.973389 | 0.180716 |
| SA 1h | 111.196 | 9.98593 | 1.05093 | 0.0694057 |
| Me-JA 1h | 75.2871 | 5.72397 | 0.902016 | 0.0945361 |
| Control 6h | 110.666 | 13.1872 | 1.0645 | 0.0728429 |
| ABA 6h | 28.6001 | 5.41183 | 0.25466 | 0.0418837 |
| ACC 6h | 113.65 | 14.2235 | 0.955282 | 0.0665836 |
| BABA 6h | 110.928 | 7.75931 | 0.966644 | 0.0674597 |
| Chitin 6h | 103.328 | 15.3855 | 0.926635 | 0.125351 |
| Epi 6h | 107.4 | 20.7887 | 0.900314 | 0.188954 |
| SA 6h | 61.7204 | 8.47026 | 0.599819 | 0.0924214 |
| Me-JA 6h | 89.6101 | 9.96244 | 0.869671 | 0.150397 |
Source Transcript PGSC0003DMT400011222 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G65600.1 | +3 | 1e-11 | 61 | 38/65 (58%) | Concanavalin A-like lectin protein kinase family protein | chr5:26216126-26218153 REVERSE LENGTH=675 |
