Probe CUST_44645_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44645_PI426222305 | JHI_St_60k_v1 | DMT400027448 | TGTGCCAATCATCATCCTTCTGTTTGGGGAGACCATTTTCTTGCCTATGCTAATCTTTCG |
All Microarray Probes Designed to Gene DMG400010587
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44645_PI426222305 | JHI_St_60k_v1 | DMT400027448 | TGTGCCAATCATCATCCTTCTGTTTGGGGAGACCATTTTCTTGCCTATGCTAATCTTTCG |
Microarray Signals from CUST_44645_PI426222305
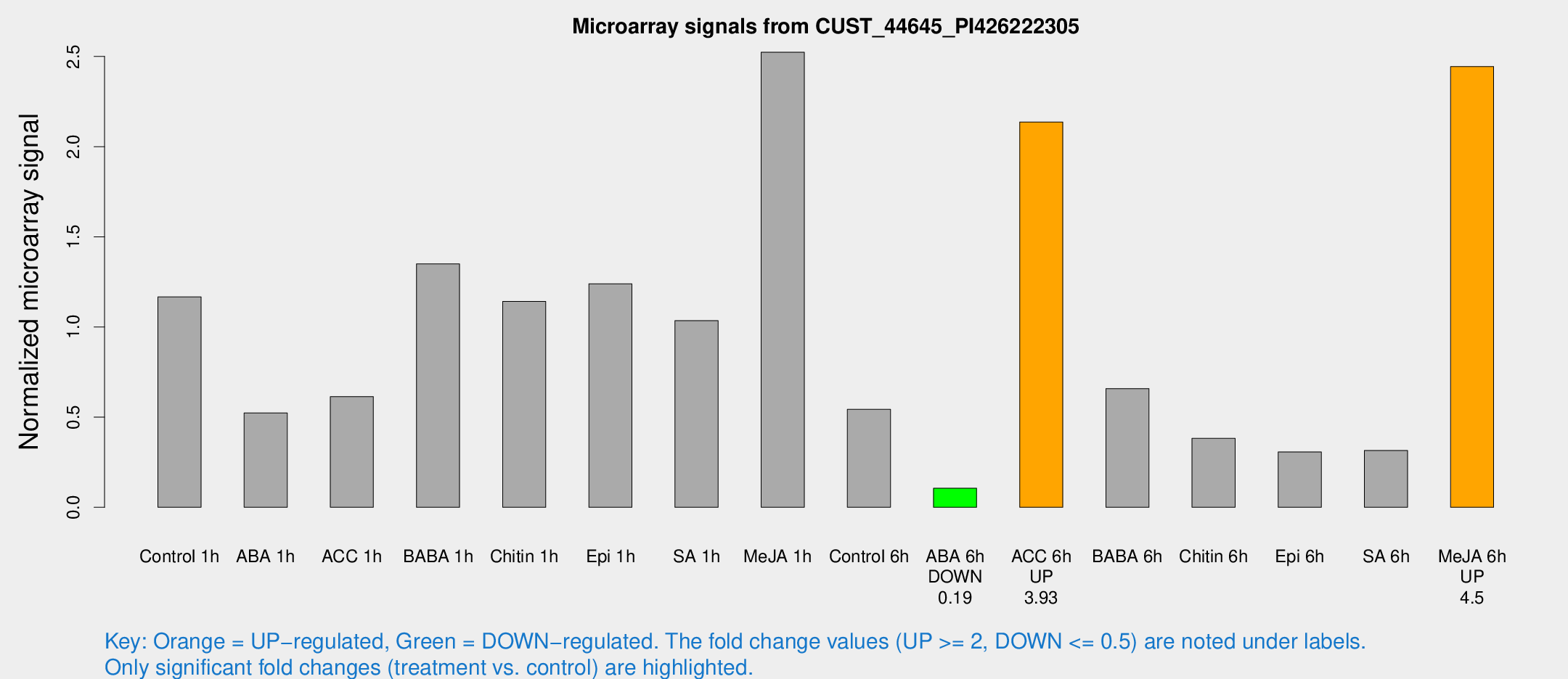
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 438.839 | 25.5788 | 1.16639 | 0.0808042 |
| ABA 1h | 189.424 | 49.5184 | 0.523292 | 0.144394 |
| ACC 1h | 277.478 | 103.909 | 0.614093 | 0.231487 |
| BABA 1h | 508.757 | 92.055 | 1.34969 | 0.133722 |
| Chitin 1h | 386.755 | 22.6314 | 1.14145 | 0.106575 |
| Epi 1h | 411.49 | 57.7805 | 1.23953 | 0.175165 |
| SA 1h | 406.018 | 51.1625 | 1.03501 | 0.127235 |
| Me-JA 1h | 809.912 | 174.995 | 2.52351 | 0.314925 |
| Control 6h | 230.183 | 82.9802 | 0.543041 | 0.177564 |
| ABA 6h | 45.4396 | 11.402 | 0.105824 | 0.0242393 |
| ACC 6h | 940.82 | 133.417 | 2.13663 | 0.62089 |
| BABA 6h | 351.716 | 176.078 | 0.658367 | 0.348167 |
| Chitin 6h | 156.309 | 23.4054 | 0.382263 | 0.0434451 |
| Epi 6h | 138.008 | 29.8389 | 0.306976 | 0.0588661 |
| SA 6h | 139.828 | 51.5589 | 0.314785 | 0.219089 |
| Me-JA 6h | 983.078 | 275.836 | 2.44391 | 0.899414 |
Source Transcript PGSC0003DMT400027448 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G23960.1 | +2 | 8e-126 | 391 | 216/547 (39%) | terpene synthase 21 | chr5:8092969-8095128 FORWARD LENGTH=547 |
