Probe CUST_44369_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44369_PI426222305 | JHI_St_60k_v1 | DMT400030955 | GCAGAGATTAGCTTTAATTGGCTTCGTTGGTGATCAGTGTCTCTGTAAAAGTTCTCATTT |
All Microarray Probes Designed to Gene DMG400011859
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44365_PI426222305 | JHI_St_60k_v1 | DMT400030954 | CCAAAGCCTTGAAGGCTAGTTCTTGATTAAATCAATCGTGAGACATGAAATTGATTGATT |
| CUST_44369_PI426222305 | JHI_St_60k_v1 | DMT400030955 | GCAGAGATTAGCTTTAATTGGCTTCGTTGGTGATCAGTGTCTCTGTAAAAGTTCTCATTT |
Microarray Signals from CUST_44369_PI426222305
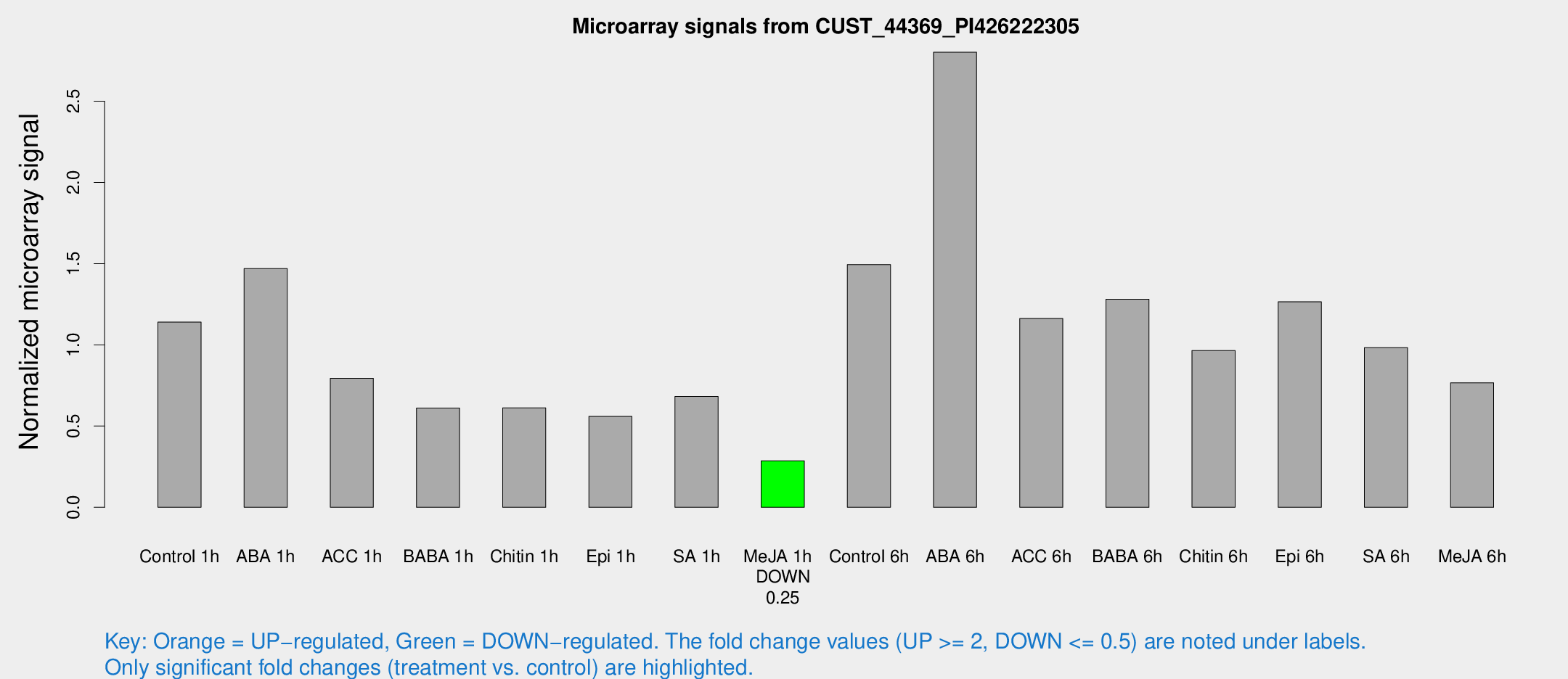
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 186.341 | 15.6195 | 1.1398 | 0.0687071 |
| ABA 1h | 220.579 | 48.2362 | 1.46974 | 0.224862 |
| ACC 1h | 133.043 | 9.51983 | 0.793855 | 0.0507007 |
| BABA 1h | 101.155 | 21.9247 | 0.611085 | 0.0887106 |
| Chitin 1h | 92.0892 | 16.6308 | 0.611916 | 0.112252 |
| Epi 1h | 84.2951 | 20.154 | 0.559495 | 0.150559 |
| SA 1h | 113.803 | 7.26983 | 0.682113 | 0.0452447 |
| Me-JA 1h | 38.5695 | 4.93907 | 0.286491 | 0.0423328 |
| Control 6h | 264.306 | 66.6927 | 1.49448 | 0.330362 |
| ABA 6h | 487.619 | 44.7447 | 2.80133 | 0.162991 |
| ACC 6h | 225.232 | 45.871 | 1.16238 | 0.0735254 |
| BABA 6h | 234.72 | 20.0309 | 1.28125 | 0.0766559 |
| Chitin 6h | 167.546 | 10.6548 | 0.965164 | 0.0902358 |
| Epi 6h | 242.326 | 50.5653 | 1.26516 | 0.340831 |
| SA 6h | 182.948 | 57.4038 | 0.982979 | 0.315655 |
| Me-JA 6h | 133.519 | 32.0309 | 0.766462 | 0.170307 |
Source Transcript PGSC0003DMT400030955 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G38480.1 | +3 | 6e-49 | 164 | 97/199 (49%) | Uncharacterised protein family (UPF0497) | chr2:16110960-16111694 REVERSE LENGTH=188 |
