Probe CUST_44024_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44024_PI426222305 | JHI_St_60k_v1 | DMT400089589 | AATTTTCAGTCAGAAGTACTTTCCGGTACCGGTGATTTTGTATCTCTCTTCCTAATATGA |
All Microarray Probes Designed to Gene DMG400039160
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44024_PI426222305 | JHI_St_60k_v1 | DMT400089589 | AATTTTCAGTCAGAAGTACTTTCCGGTACCGGTGATTTTGTATCTCTCTTCCTAATATGA |
Microarray Signals from CUST_44024_PI426222305
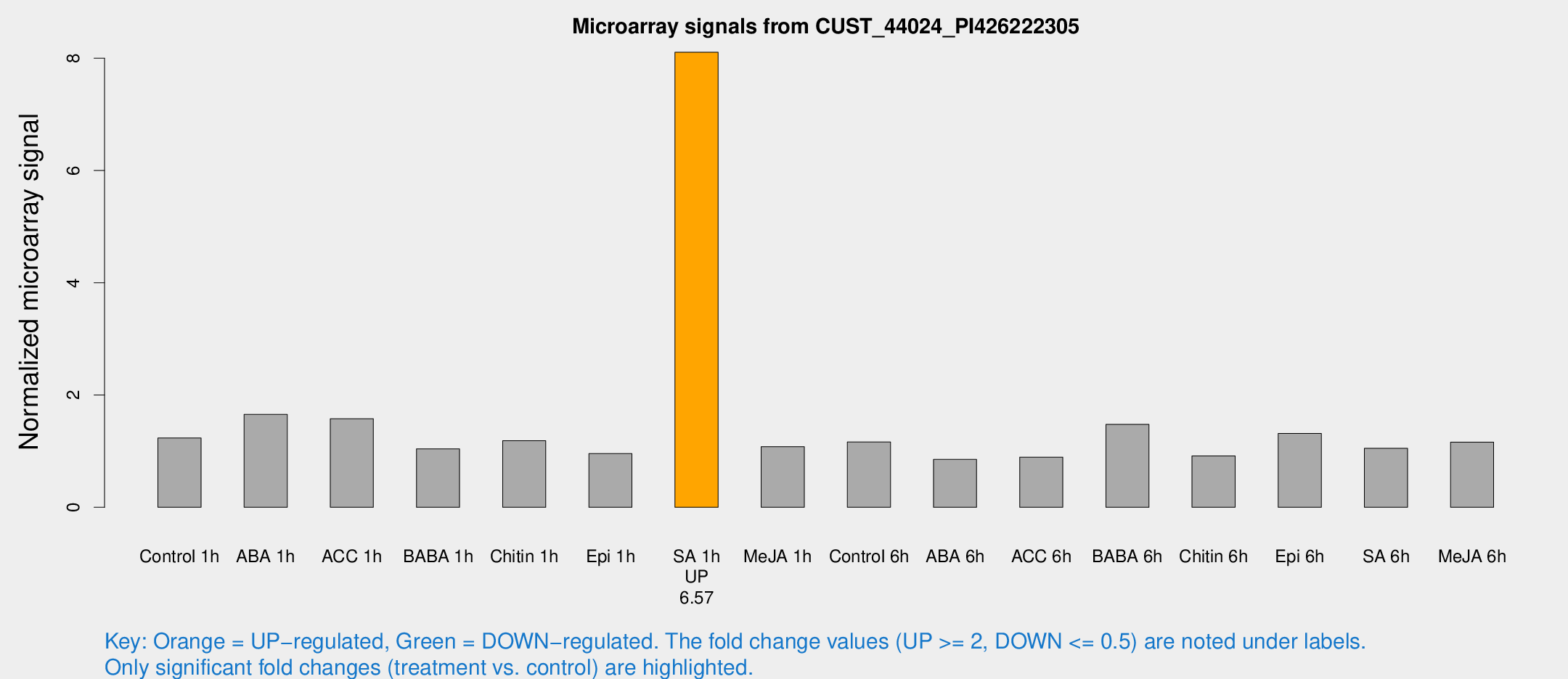
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.80854 | 3.6948 | 1.23404 | 0.567865 |
| ABA 1h | 11.211 | 3.40897 | 1.65424 | 0.616687 |
| ACC 1h | 12.9578 | 4.82749 | 1.57713 | 0.706347 |
| BABA 1h | 7.06869 | 3.6798 | 1.04012 | 0.550775 |
| Chitin 1h | 7.8098 | 3.46031 | 1.18738 | 0.560111 |
| Epi 1h | 5.79794 | 3.36931 | 0.955893 | 0.554384 |
| SA 1h | 58.8576 | 5.8396 | 8.10786 | 1.04559 |
| Me-JA 1h | 6.1898 | 3.60007 | 1.08143 | 0.627032 |
| Control 6h | 8.72206 | 3.60308 | 1.16424 | 0.550923 |
| ABA 6h | 6.34316 | 3.67671 | 0.85274 | 0.493951 |
| ACC 6h | 7.33962 | 4.3593 | 0.893308 | 0.517288 |
| BABA 6h | 12.513 | 4.04771 | 1.47715 | 0.564982 |
| Chitin 6h | 6.82428 | 3.95192 | 0.916825 | 0.530886 |
| Epi 6h | 11.67 | 4.36133 | 1.31506 | 0.615896 |
| SA 6h | 7.29828 | 3.84043 | 1.05013 | 0.559721 |
| Me-JA 6h | 8.72876 | 3.61645 | 1.16111 | 0.559287 |
Source Transcript PGSC0003DMT400089589 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G22910.1 | +1 | 4e-112 | 368 | 185/248 (75%) | ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein | chr3:8116335-8119388 REVERSE LENGTH=1017 |
