Probe CUST_44023_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44023_PI426222305 | JHI_St_60k_v1 | DMT400089336 | CCCAGAGAGACCATTCTTTAGTTATCTAAGGTTGAATTACTTGAAAGCCTTGTTTAAATG |
All Microarray Probes Designed to Gene DMG400038907
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44023_PI426222305 | JHI_St_60k_v1 | DMT400089336 | CCCAGAGAGACCATTCTTTAGTTATCTAAGGTTGAATTACTTGAAAGCCTTGTTTAAATG |
Microarray Signals from CUST_44023_PI426222305
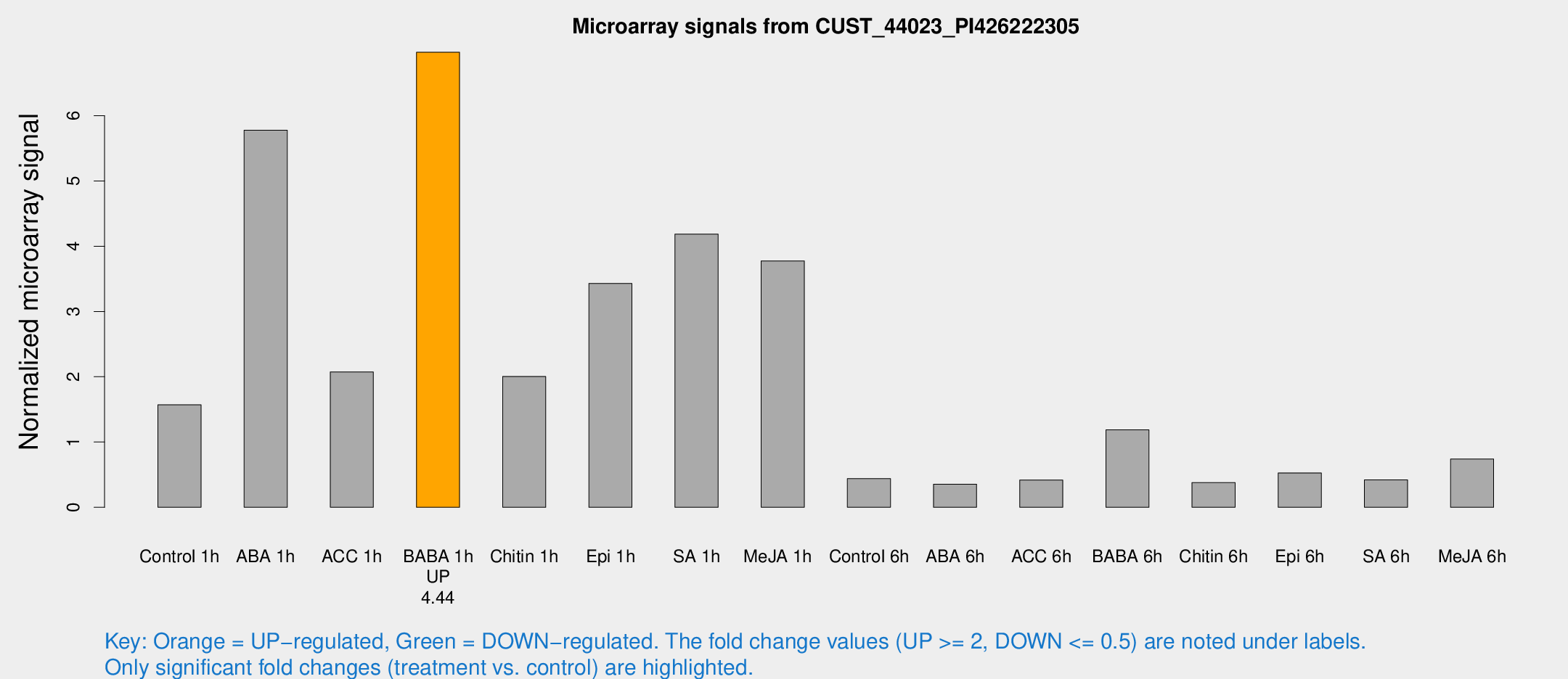
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 39.0039 | 14.3373 | 1.57042 | 0.703834 |
| ABA 1h | 112.465 | 8.0732 | 5.77589 | 0.365299 |
| ACC 1h | 47.5482 | 6.27164 | 2.07465 | 0.203802 |
| BABA 1h | 152.33 | 27.0362 | 6.97167 | 2.05922 |
| Chitin 1h | 43.028 | 11.7452 | 2.00434 | 0.804122 |
| Epi 1h | 70.1511 | 18.3466 | 3.42992 | 1.14287 |
| SA 1h | 98.9084 | 20.5439 | 4.18605 | 0.723185 |
| Me-JA 1h | 68.7936 | 9.06878 | 3.77519 | 0.278836 |
| Control 6h | 10.6749 | 3.22075 | 0.439697 | 0.163206 |
| ABA 6h | 10.1296 | 4.80579 | 0.3539 | 0.181991 |
| ACC 6h | 11.1983 | 3.80704 | 0.418005 | 0.160734 |
| BABA 6h | 35.7141 | 16.0112 | 1.18624 | 0.593528 |
| Chitin 6h | 9.28747 | 3.41541 | 0.378238 | 0.156865 |
| Epi 6h | 14.831 | 5.31975 | 0.526571 | 0.236643 |
| SA 6h | 10.2962 | 3.78382 | 0.419775 | 0.172751 |
| Me-JA 6h | 16.2514 | 3.19302 | 0.739604 | 0.147921 |
Source Transcript PGSC0003DMT400089336 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G22910.1 | +1 | 0.0 | 850 | 487/771 (63%) | ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein | chr3:8116335-8119388 REVERSE LENGTH=1017 |
