Probe CUST_44018_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44018_PI426222305 | JHI_St_60k_v1 | DMT400096782 | AACAGAATGTGGGATTCTCCATCCTAATCCGGAAGTAGATGAAGGAGCAGGTCATAGAAG |
All Microarray Probes Designed to Gene DMG400046353
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44018_PI426222305 | JHI_St_60k_v1 | DMT400096782 | AACAGAATGTGGGATTCTCCATCCTAATCCGGAAGTAGATGAAGGAGCAGGTCATAGAAG |
Microarray Signals from CUST_44018_PI426222305
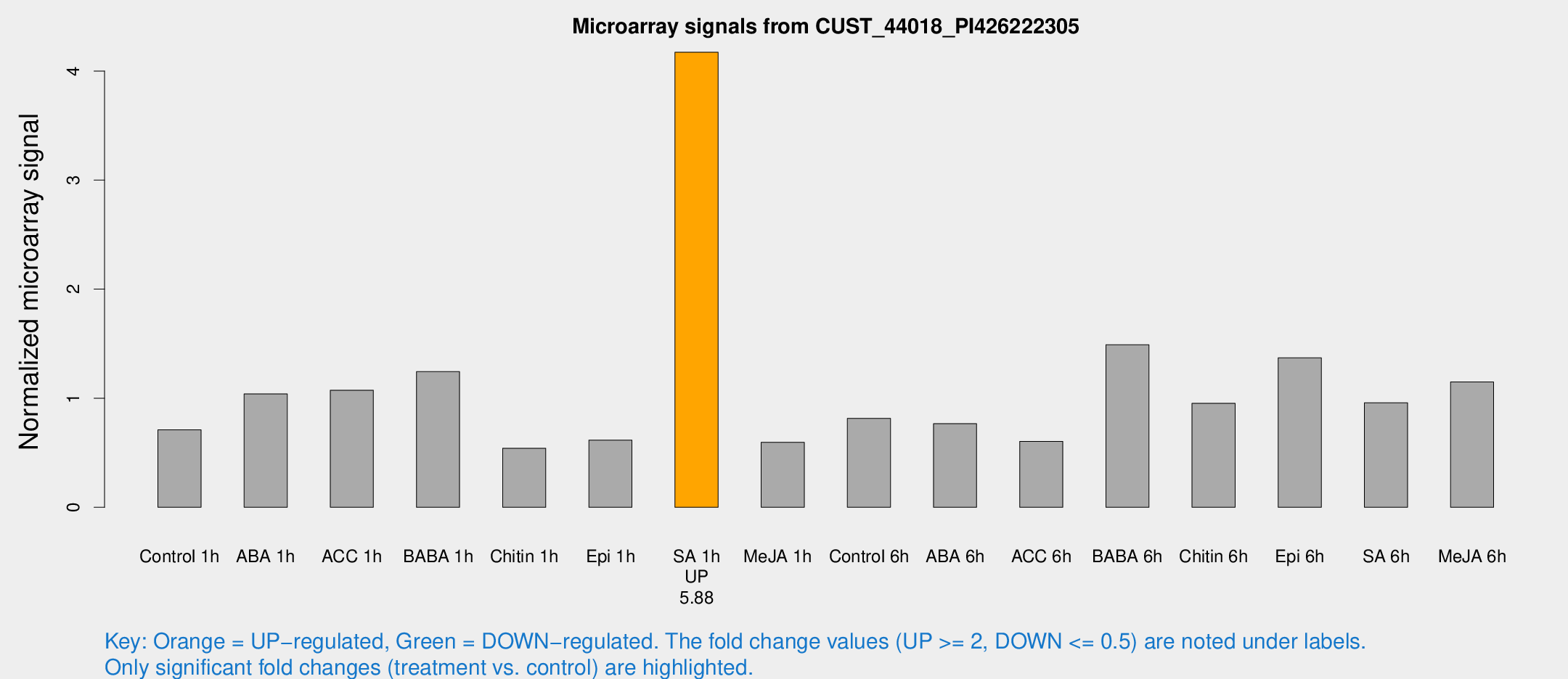
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.5346 | 5.73527 | 0.709958 | 0.324672 |
| ABA 1h | 13.9325 | 4.58403 | 1.03926 | 0.513685 |
| ACC 1h | 15.9743 | 4.85902 | 1.07374 | 0.44427 |
| BABA 1h | 15.5242 | 3.61992 | 1.24348 | 0.291291 |
| Chitin 1h | 6.38486 | 3.38249 | 0.541712 | 0.290772 |
| Epi 1h | 7.28122 | 3.36048 | 0.616366 | 0.309937 |
| SA 1h | 56.5521 | 7.70194 | 4.17224 | 0.733436 |
| Me-JA 1h | 6.30738 | 3.42204 | 0.596464 | 0.324888 |
| Control 6h | 10.6241 | 3.51678 | 0.815439 | 0.273687 |
| ABA 6h | 11.347 | 3.64084 | 0.766356 | 0.294916 |
| ACC 6h | 9.4884 | 4.27535 | 0.604444 | 0.292329 |
| BABA 6h | 21.7205 | 4.067 | 1.49018 | 0.280452 |
| Chitin 6h | 13.4225 | 3.8481 | 0.95357 | 0.285616 |
| Epi 6h | 22.1412 | 6.79151 | 1.37059 | 0.562064 |
| SA 6h | 14.72 | 6.20219 | 0.958685 | 0.479768 |
| Me-JA 6h | 19.3463 | 9.15186 | 1.14911 | 0.620605 |
Source Transcript PGSC0003DMT400096782 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G63380.1 | +1 | 0.0 | 717 | 403/634 (64%) | ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein | chr3:23407112-23410213 REVERSE LENGTH=1033 |
