Probe CUST_43992_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43992_PI426222305 | JHI_St_60k_v1 | DMT400093936 | AAGTTTGCAAATACAGAGAGGCTGAATTGGGGGCAATGGGGAATCTGCATTGGATTTGCA |
All Microarray Probes Designed to Gene DMG400043507
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43992_PI426222305 | JHI_St_60k_v1 | DMT400093936 | AAGTTTGCAAATACAGAGAGGCTGAATTGGGGGCAATGGGGAATCTGCATTGGATTTGCA |
Microarray Signals from CUST_43992_PI426222305
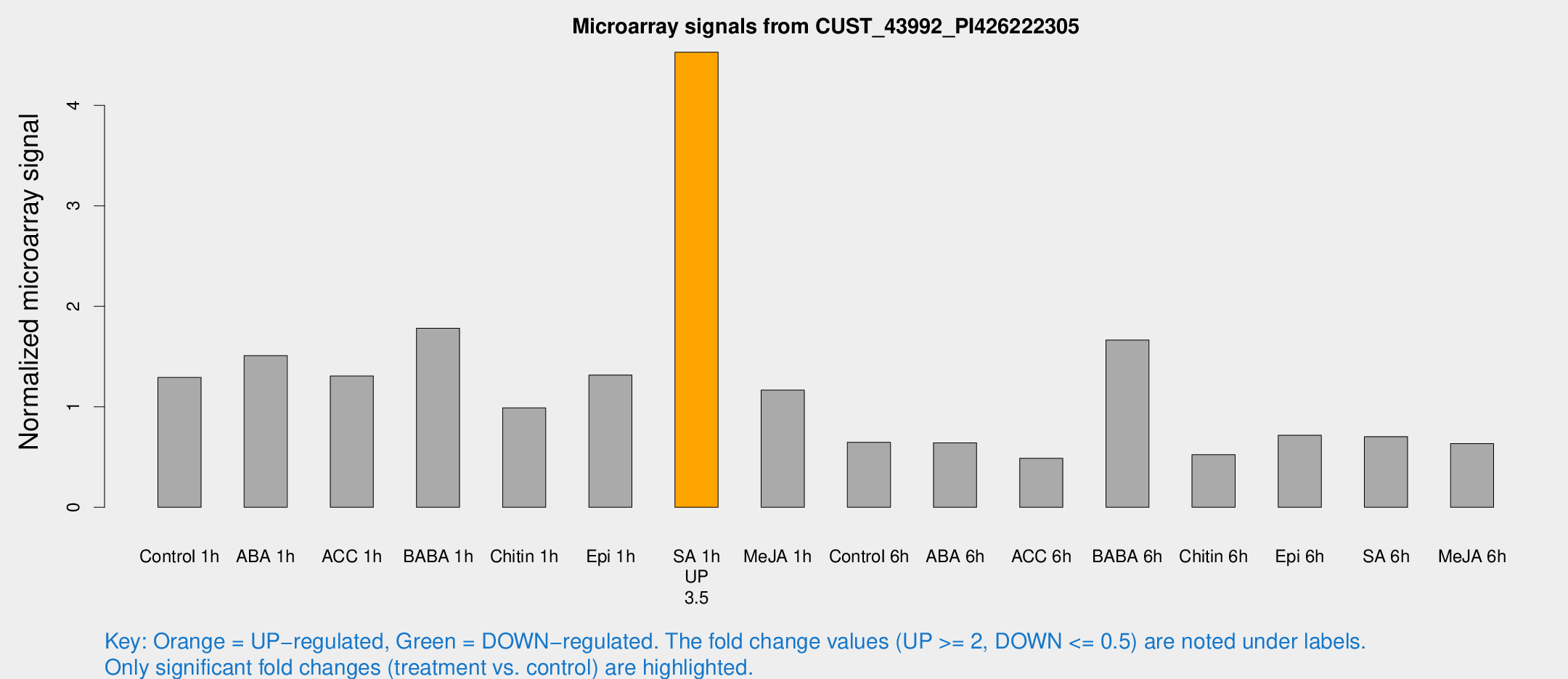
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 694.134 | 171.003 | 1.2923 | 0.279327 |
| ABA 1h | 677.226 | 39.2825 | 1.5102 | 0.0874393 |
| ACC 1h | 739.002 | 187.044 | 1.30717 | 0.311533 |
| BABA 1h | 874.08 | 63.9294 | 1.78216 | 0.309406 |
| Chitin 1h | 452.8 | 30.7387 | 0.990425 | 0.0644344 |
| Epi 1h | 582.416 | 61.7151 | 1.31571 | 0.100012 |
| SA 1h | 2358.02 | 136.395 | 4.52861 | 0.261526 |
| Me-JA 1h | 481.889 | 28.0024 | 1.16626 | 0.0961754 |
| Control 6h | 377.182 | 121.72 | 0.647448 | 0.209944 |
| ABA 6h | 349.478 | 36.346 | 0.642094 | 0.0717663 |
| ACC 6h | 283.861 | 16.8842 | 0.487705 | 0.0802597 |
| BABA 6h | 949.996 | 77.3933 | 1.66531 | 0.164815 |
| Chitin 6h | 288.805 | 45.9458 | 0.523693 | 0.0809583 |
| Epi 6h | 431.142 | 102.487 | 0.716909 | 0.184554 |
| SA 6h | 373.887 | 82.709 | 0.702888 | 0.0993565 |
| Me-JA 6h | 338.898 | 84.8876 | 0.634374 | 0.126341 |
Source Transcript PGSC0003DMT400093936 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G22910.1 | +1 | 0.0 | 741 | 381/550 (69%) | ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein | chr3:8116335-8119388 REVERSE LENGTH=1017 |
