Probe CUST_43929_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43929_PI426222305 | JHI_St_60k_v1 | DMT400044114 | TTTGGGTTTGCACATGGGGCATTAATAGTCCCTTATGTTTGTAATATTGGCGTACTTTTA |
All Microarray Probes Designed to Gene DMG400017119
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43929_PI426222305 | JHI_St_60k_v1 | DMT400044114 | TTTGGGTTTGCACATGGGGCATTAATAGTCCCTTATGTTTGTAATATTGGCGTACTTTTA |
Microarray Signals from CUST_43929_PI426222305
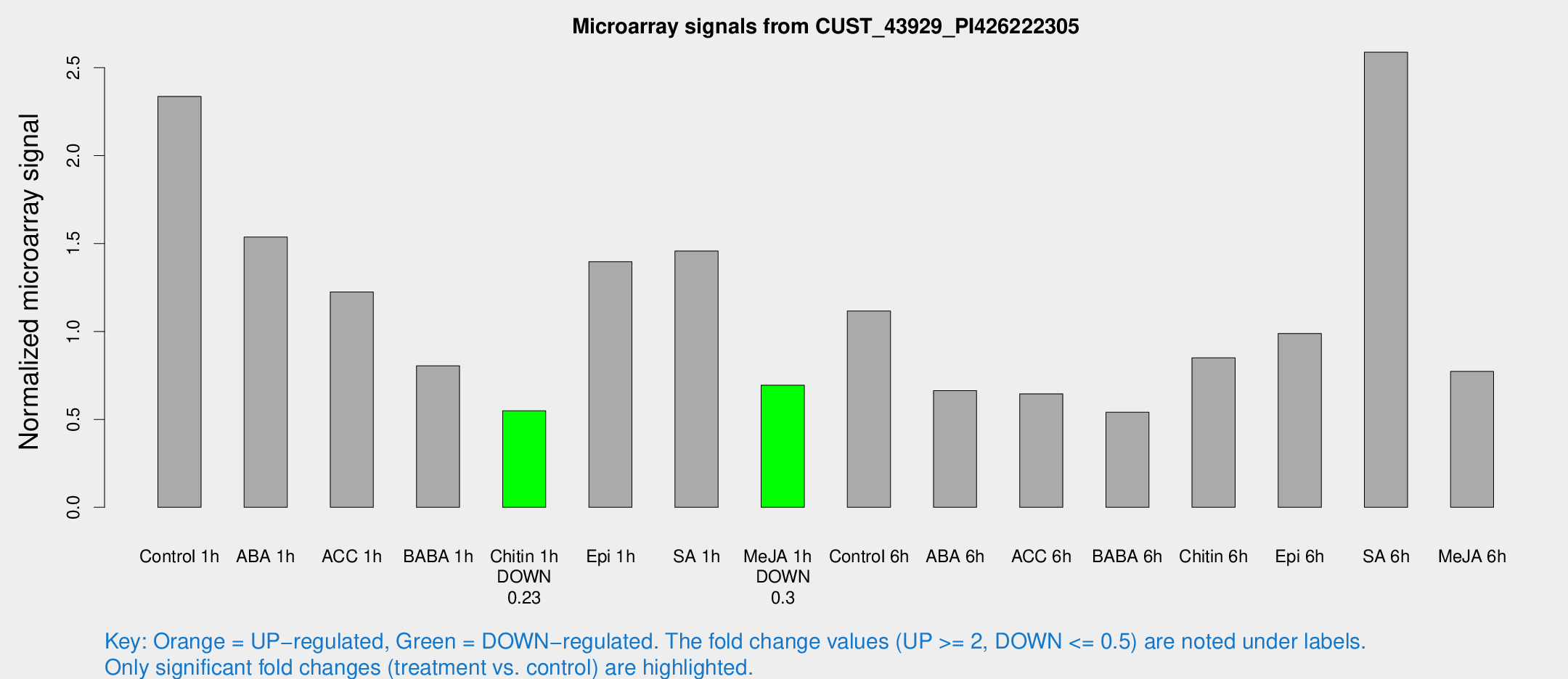
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 829.147 | 86.1063 | 2.33665 | 0.135336 |
| ABA 1h | 503.692 | 122.242 | 1.53676 | 0.399574 |
| ACC 1h | 481.289 | 128.13 | 1.22478 | 0.300226 |
| BABA 1h | 311.349 | 94.4369 | 0.804749 | 0.250405 |
| Chitin 1h | 173.952 | 11.6004 | 0.548564 | 0.072597 |
| Epi 1h | 426.875 | 32.3097 | 1.39689 | 0.11245 |
| SA 1h | 534.993 | 70.0622 | 1.45746 | 0.103626 |
| Me-JA 1h | 205.994 | 39.8702 | 0.69435 | 0.0767907 |
| Control 6h | 426.081 | 126.928 | 1.1166 | 0.321724 |
| ABA 6h | 247.783 | 14.8618 | 0.663206 | 0.0397402 |
| ACC 6h | 265.781 | 37.2988 | 0.645432 | 0.0920565 |
| BABA 6h | 243.553 | 86.6556 | 0.540619 | 0.238517 |
| Chitin 6h | 329.156 | 64.1096 | 0.849864 | 0.141362 |
| Epi 6h | 398.244 | 50.0514 | 0.988797 | 0.0833547 |
| SA 6h | 936.811 | 174.509 | 2.58816 | 0.238156 |
| Me-JA 6h | 280.882 | 57.6178 | 0.772471 | 0.119556 |
Source Transcript PGSC0003DMT400044114 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G36780.1 | +2 | 4e-170 | 499 | 245/417 (59%) | UDP-Glycosyltransferase superfamily protein | chr2:15417618-15419108 REVERSE LENGTH=496 |
