Probe CUST_43833_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43833_PI426222305 | JHI_St_60k_v1 | DMT400040206 | AATGTTGATTTCAAGTTATGGCCAACAACACCAATTGCATCTCCTGTTTTGCGATGCTTT |
All Microarray Probes Designed to Gene DMG400015557
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43833_PI426222305 | JHI_St_60k_v1 | DMT400040206 | AATGTTGATTTCAAGTTATGGCCAACAACACCAATTGCATCTCCTGTTTTGCGATGCTTT |
Microarray Signals from CUST_43833_PI426222305
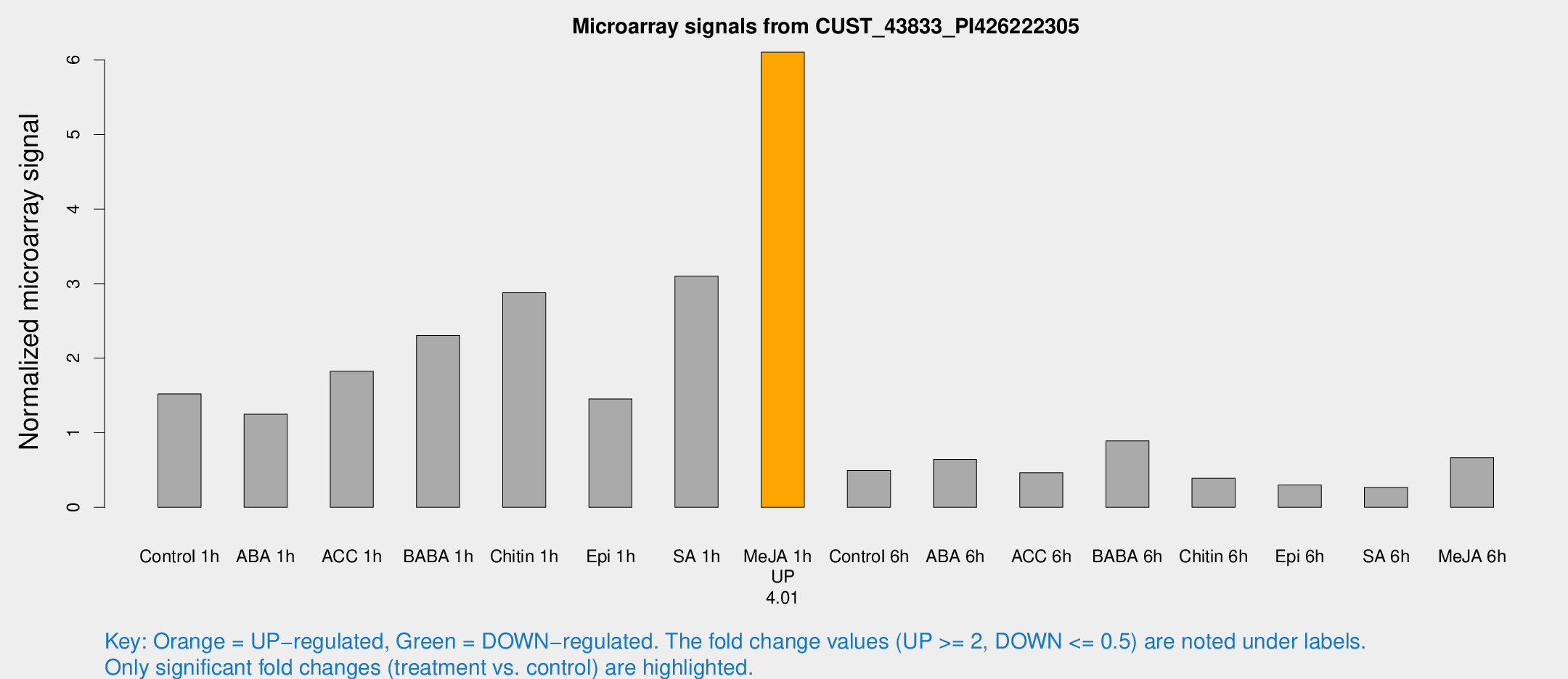
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 487.066 | 76.9716 | 1.52204 | 0.229997 |
| ABA 1h | 388.928 | 143.051 | 1.24847 | 0.559407 |
| ACC 1h | 647.853 | 190.959 | 1.82475 | 0.495629 |
| BABA 1h | 733.827 | 174.934 | 2.3052 | 0.373672 |
| Chitin 1h | 815.037 | 81.4591 | 2.87892 | 0.448206 |
| Epi 1h | 399.519 | 51.9822 | 1.45299 | 0.198306 |
| SA 1h | 1046.83 | 230.619 | 3.09899 | 0.64538 |
| Me-JA 1h | 1566.27 | 142.022 | 6.10367 | 0.543001 |
| Control 6h | 203.653 | 105.443 | 0.495759 | 0.254615 |
| ABA 6h | 220.791 | 41.2792 | 0.639422 | 0.105273 |
| ACC 6h | 167.342 | 14.9529 | 0.46398 | 0.0569313 |
| BABA 6h | 325.761 | 72.3054 | 0.891107 | 0.160864 |
| Chitin 6h | 132.566 | 21.4021 | 0.389191 | 0.0716338 |
| Epi 6h | 107.364 | 13.3613 | 0.301021 | 0.0373605 |
| SA 6h | 81.9498 | 5.86553 | 0.264918 | 0.0283076 |
| Me-JA 6h | 216.156 | 46.3125 | 0.667757 | 0.0937773 |
Source Transcript PGSC0003DMT400040206 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37430.1 | +2 | 7e-34 | 122 | 73/163 (45%) | C2H2 and C2HC zinc fingers superfamily protein | chr2:15706454-15706990 FORWARD LENGTH=178 |
