Probe CUST_43788_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43788_PI426222305 | JHI_St_60k_v1 | DMT400040141 | GGCCGATGGCACCAATTGCATCTCCTGTTTTGAGAACTTTTATTTAAGTCGTTGTTAAAC |
All Microarray Probes Designed to Gene DMG400015531
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43788_PI426222305 | JHI_St_60k_v1 | DMT400040141 | GGCCGATGGCACCAATTGCATCTCCTGTTTTGAGAACTTTTATTTAAGTCGTTGTTAAAC |
Microarray Signals from CUST_43788_PI426222305
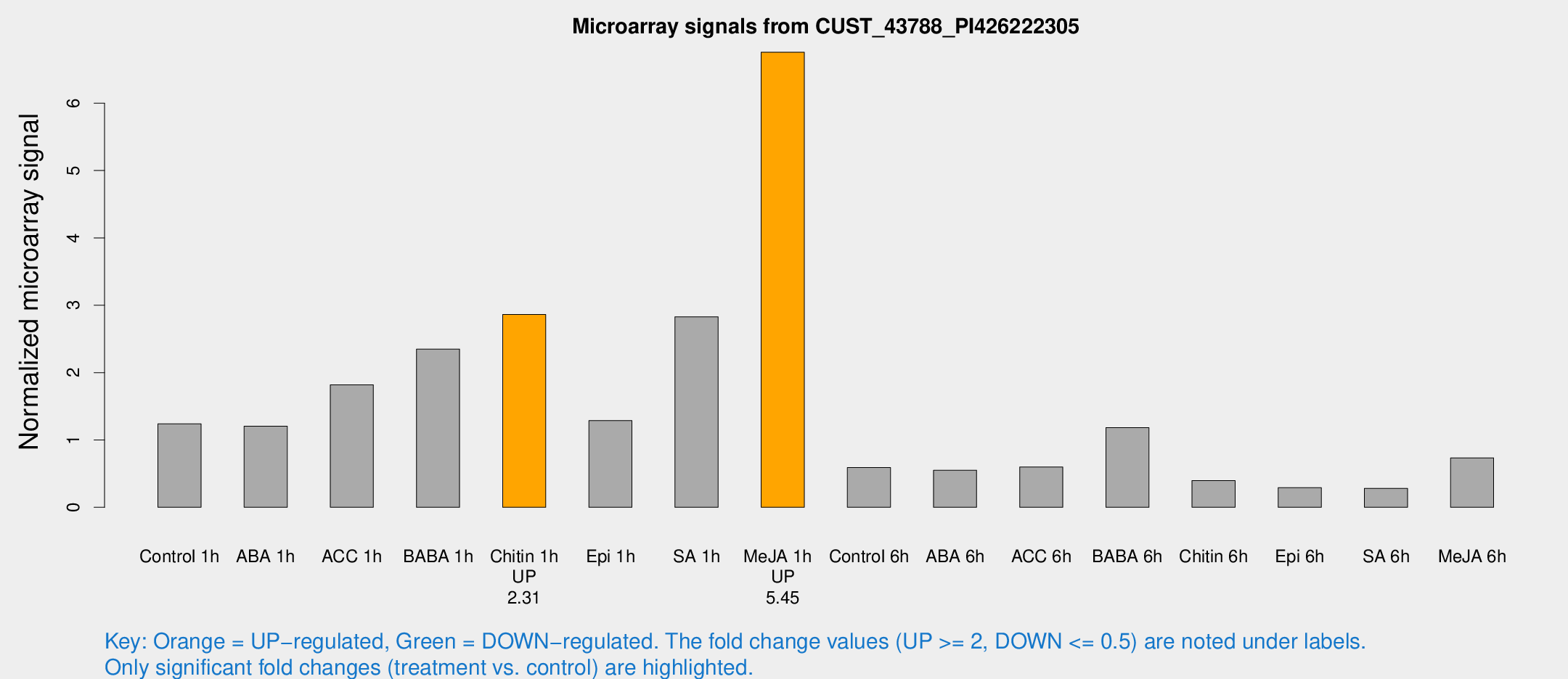
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 271.43 | 59.4814 | 1.23905 | 0.266194 |
| ABA 1h | 263.016 | 113.675 | 1.20467 | 0.64574 |
| ACC 1h | 420.413 | 109.101 | 1.81808 | 0.400804 |
| BABA 1h | 501.643 | 125.368 | 2.34882 | 0.409341 |
| Chitin 1h | 538.629 | 39.002 | 2.86259 | 0.367862 |
| Epi 1h | 235.865 | 29.3688 | 1.28699 | 0.159767 |
| SA 1h | 645.41 | 149.82 | 2.82754 | 0.654641 |
| Me-JA 1h | 1153.17 | 76.5725 | 6.75652 | 0.691318 |
| Control 6h | 162.399 | 87.4037 | 0.58971 | 0.305008 |
| ABA 6h | 123.789 | 15.4847 | 0.549988 | 0.0446462 |
| ACC 6h | 149.507 | 29.0961 | 0.59815 | 0.127292 |
| BABA 6h | 284.535 | 51.6287 | 1.18282 | 0.195311 |
| Chitin 6h | 88.869 | 8.83569 | 0.397281 | 0.0504341 |
| Epi 6h | 68.761 | 5.27435 | 0.291667 | 0.0400047 |
| SA 6h | 57.9958 | 4.60561 | 0.281344 | 0.0361447 |
| Me-JA 6h | 160.464 | 38.4852 | 0.733542 | 0.113112 |
Source Transcript PGSC0003DMT400040141 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37430.1 | +1 | 2e-35 | 126 | 75/164 (46%) | C2H2 and C2HC zinc fingers superfamily protein | chr2:15706454-15706990 FORWARD LENGTH=178 |
