Probe CUST_43706_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43706_PI426222305 | JHI_St_60k_v1 | DMT400004843 | TGGGTTCATGGGTTTCTTTTGGTCGTTGGATTATTCTTTGCGGCGTTTATTGAAATTTAG |
All Microarray Probes Designed to Gene DMG400001923
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43673_PI426222305 | JHI_St_60k_v1 | DMT400004844 | GGTTTCTTTTGGTCGTTGGATTATTCTTTGCGGCGTTTATTGAAATTTAGTTGATGTGAT |
| CUST_43706_PI426222305 | JHI_St_60k_v1 | DMT400004843 | TGGGTTCATGGGTTTCTTTTGGTCGTTGGATTATTCTTTGCGGCGTTTATTGAAATTTAG |
Microarray Signals from CUST_43706_PI426222305
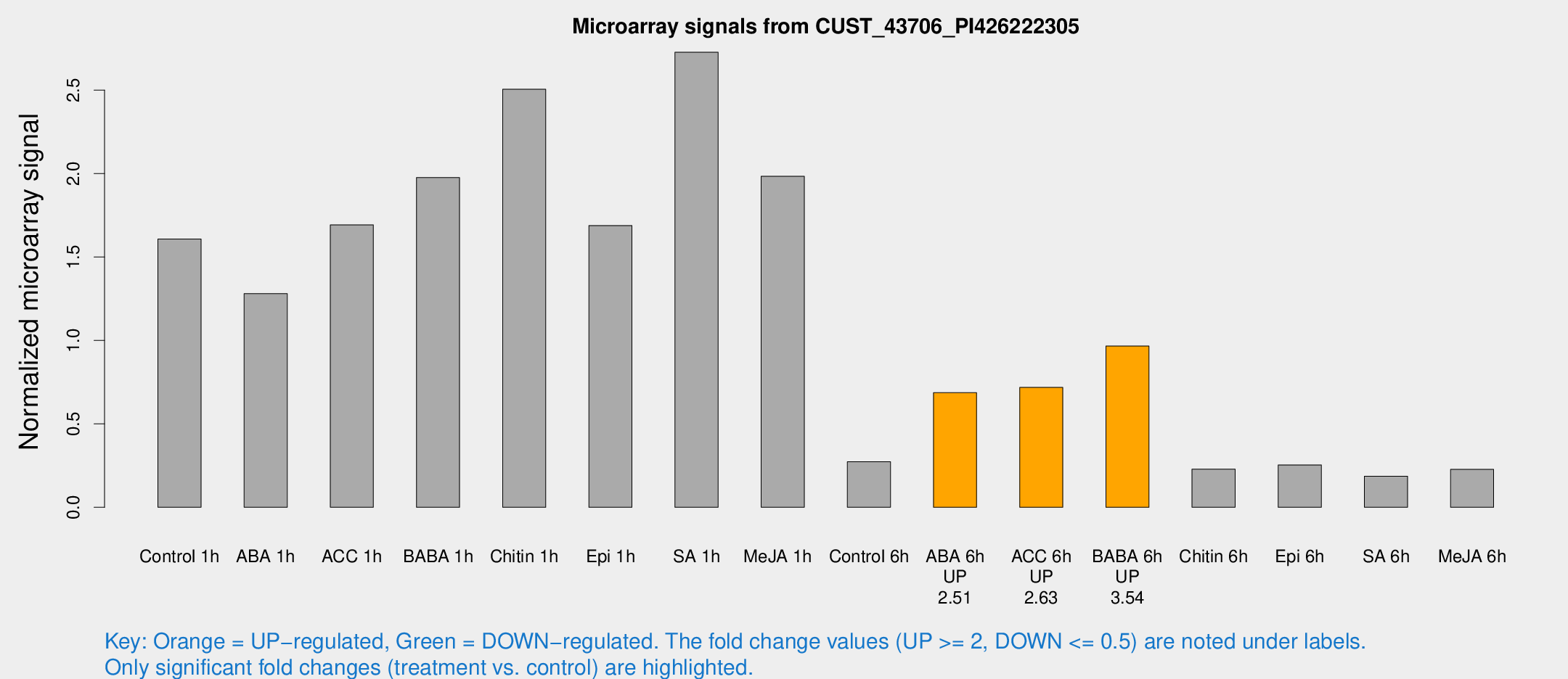
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15747.2 | 2221.63 | 1.60755 | 0.117541 |
| ABA 1h | 11268 | 1985.59 | 1.28047 | 0.287975 |
| ACC 1h | 18221.1 | 4710.77 | 1.69211 | 0.414257 |
| BABA 1h | 19433.4 | 4565.1 | 1.97552 | 0.317392 |
| Chitin 1h | 21731 | 1452.76 | 2.50471 | 0.212872 |
| Epi 1h | 14098.9 | 932.264 | 1.68855 | 0.0974892 |
| SA 1h | 27304.4 | 3175.68 | 2.72692 | 0.237893 |
| Me-JA 1h | 15745.3 | 1686.79 | 1.98362 | 0.114526 |
| Control 6h | 2741.94 | 521.765 | 0.273282 | 0.0313381 |
| ABA 6h | 7511.65 | 1844.43 | 0.68676 | 0.153993 |
| ACC 6h | 7998.18 | 779.927 | 0.718443 | 0.0864636 |
| BABA 6h | 11540 | 3894.28 | 0.966075 | 0.302867 |
| Chitin 6h | 2348.38 | 135.972 | 0.229069 | 0.013231 |
| Epi 6h | 2804.22 | 383.541 | 0.254027 | 0.0281316 |
| SA 6h | 1906.27 | 469.009 | 0.186028 | 0.0325622 |
| Me-JA 6h | 2280.03 | 505.457 | 0.227119 | 0.0313892 |
Source Transcript PGSC0003DMT400004843 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G24140.1 | +1 | 6e-134 | 392 | 200/358 (56%) | Matrixin family protein | chr1:8536131-8537285 REVERSE LENGTH=384 |
