Probe CUST_43657_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43657_PI426222305 | JHI_St_60k_v1 | DMT400030787 | CACACGGACATGAAGATCAGAATGAAGTTTAAGTTTGGATTAATCTTCATTAGTACCCAA |
All Microarray Probes Designed to Gene DMG400011793
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43657_PI426222305 | JHI_St_60k_v1 | DMT400030787 | CACACGGACATGAAGATCAGAATGAAGTTTAAGTTTGGATTAATCTTCATTAGTACCCAA |
| CUST_43627_PI426222305 | JHI_St_60k_v1 | DMT400030786 | GTGCTAGATTGGACCATCGGAAATCTTACTTGTGTTGAAGCAAAGATTATGCTTGTCTAG |
Microarray Signals from CUST_43657_PI426222305
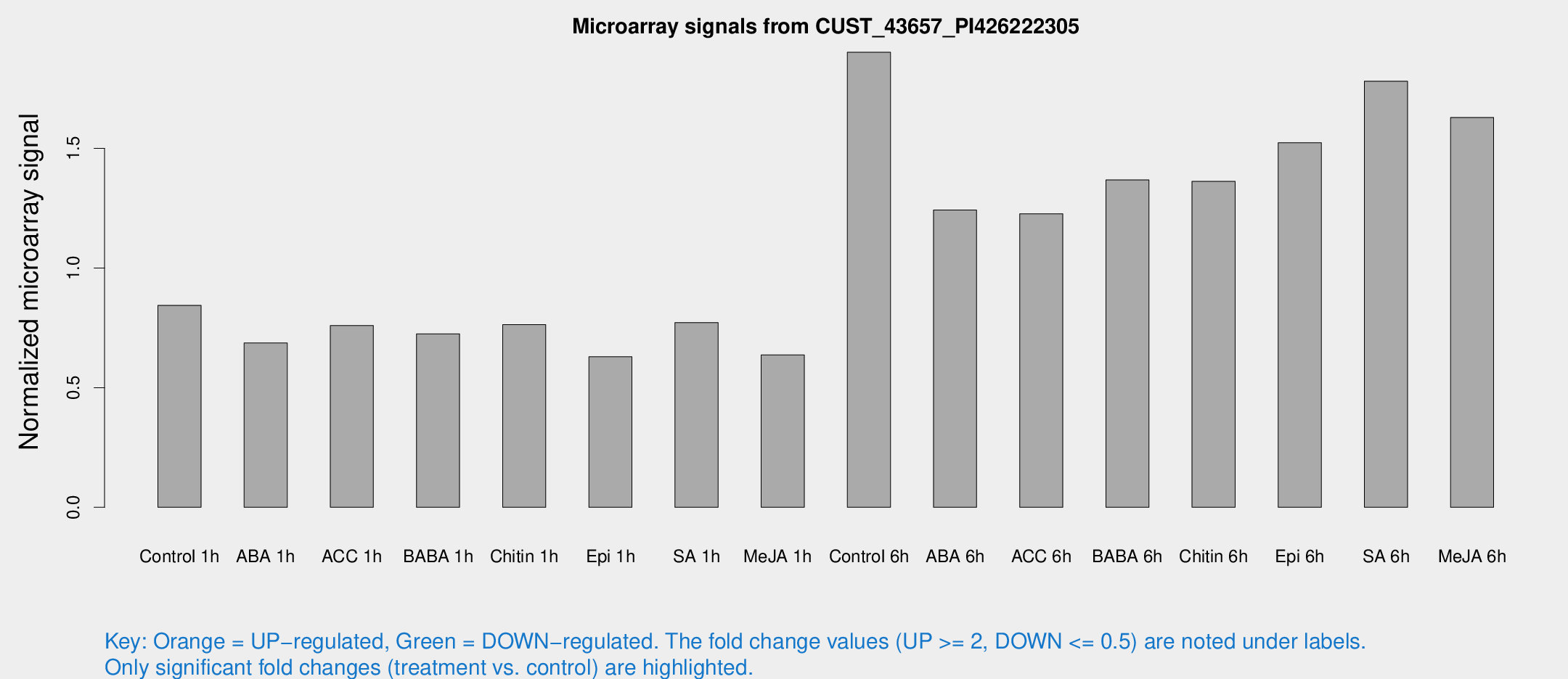
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 150.967 | 27.4764 | 0.844018 | 0.108969 |
| ABA 1h | 105.516 | 8.34193 | 0.686677 | 0.0831567 |
| ACC 1h | 139.996 | 27.0646 | 0.759382 | 0.0980723 |
| BABA 1h | 129.311 | 31.1061 | 0.724858 | 0.12872 |
| Chitin 1h | 119.778 | 12.6344 | 0.763085 | 0.0697799 |
| Epi 1h | 94.8679 | 9.54419 | 0.628928 | 0.0453701 |
| SA 1h | 138.91 | 16.76 | 0.771883 | 0.0492275 |
| Me-JA 1h | 91.5173 | 12.7054 | 0.63692 | 0.0456791 |
| Control 6h | 345.407 | 70.1157 | 1.90156 | 0.273531 |
| ABA 6h | 230.462 | 24.6832 | 1.24212 | 0.0958997 |
| ACC 6h | 245.534 | 25.1346 | 1.22658 | 0.0931144 |
| BABA 6h | 264.391 | 15.8354 | 1.36785 | 0.0818687 |
| Chitin 6h | 251.254 | 16.5852 | 1.36169 | 0.118745 |
| Epi 6h | 303.494 | 43.933 | 1.52341 | 0.345102 |
| SA 6h | 318.882 | 63.6093 | 1.78061 | 0.195259 |
| Me-JA 6h | 295.198 | 68.3992 | 1.62885 | 0.243008 |
Source Transcript PGSC0003DMT400030787 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G21270.1 | +1 | 1e-147 | 418 | 231/458 (50%) | wall-associated kinase 2 | chr1:7444997-7447345 FORWARD LENGTH=732 |
