Probe CUST_43323_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43323_PI426222305 | JHI_St_60k_v1 | DMT400064455 | TACCGGCTCAAATAACTCGGCAGTAAAAGCATTTACTACAACTTCAGCTGTTTTTGTGTC |
All Microarray Probes Designed to Gene DMG400025042
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43302_PI426222305 | JHI_St_60k_v1 | DMT400064456 | TGACACTCTTGTGTTCAAGTACATGAAGGGATCAAACTCAGTATTAGTGGTAAACAAGGA |
| CUST_43323_PI426222305 | JHI_St_60k_v1 | DMT400064455 | TACCGGCTCAAATAACTCGGCAGTAAAAGCATTTACTACAACTTCAGCTGTTTTTGTGTC |
Microarray Signals from CUST_43323_PI426222305
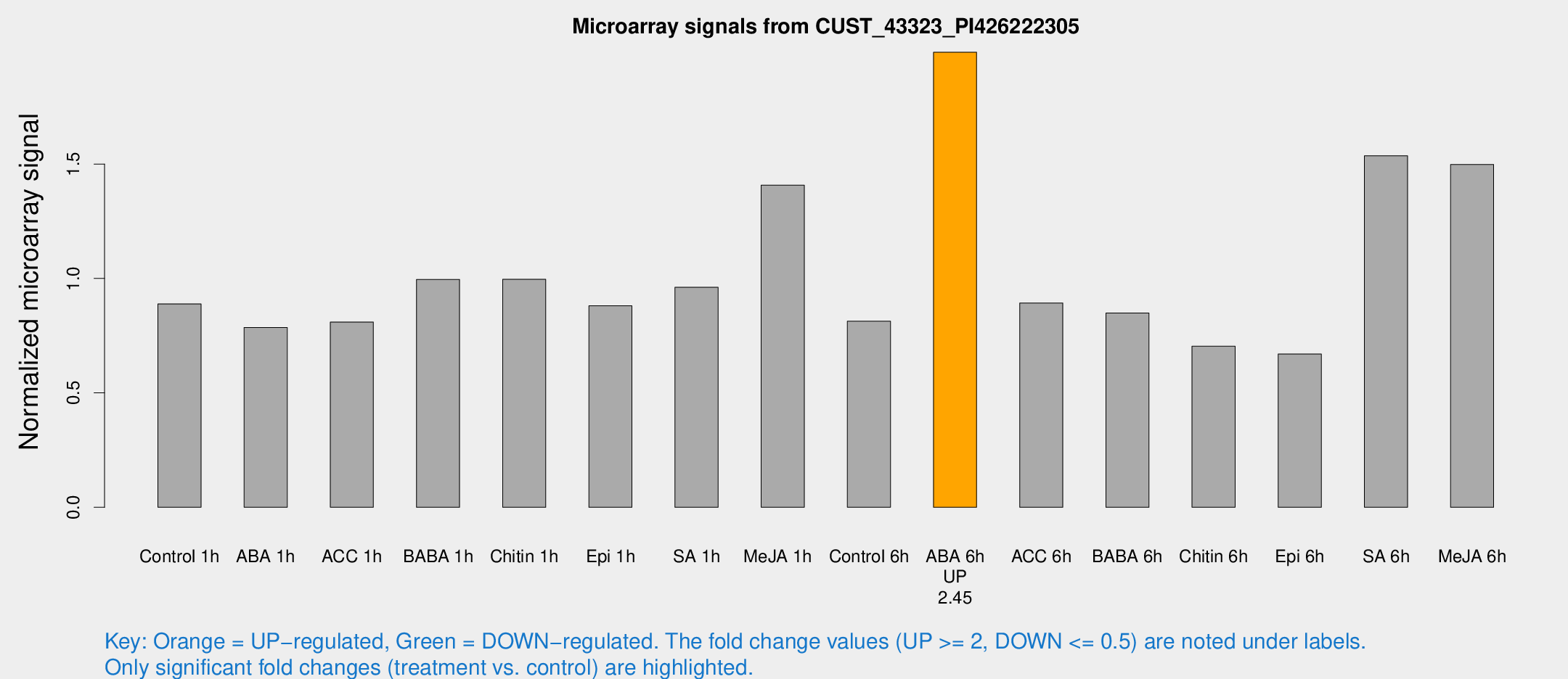
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6006.94 | 703.633 | 0.888786 | 0.0513161 |
| ABA 1h | 4870.66 | 1028.56 | 0.785963 | 0.156785 |
| ACC 1h | 6026.04 | 1722.39 | 0.809538 | 0.196542 |
| BABA 1h | 6970.96 | 1781.72 | 0.995732 | 0.209267 |
| Chitin 1h | 6170.65 | 1133.59 | 0.99611 | 0.100887 |
| Epi 1h | 5216.26 | 797.541 | 0.880549 | 0.138739 |
| SA 1h | 7068.63 | 1662.87 | 0.961273 | 0.224386 |
| Me-JA 1h | 7972.76 | 1460.5 | 1.40806 | 0.179122 |
| Control 6h | 5632.67 | 1043.9 | 0.813287 | 0.10427 |
| ABA 6h | 14968.7 | 3415.13 | 1.98862 | 0.406827 |
| ACC 6h | 6872.58 | 505.023 | 0.89294 | 0.193229 |
| BABA 6h | 6444.31 | 842.027 | 0.848666 | 0.0995701 |
| Chitin 6h | 5011.84 | 290.223 | 0.703598 | 0.0406251 |
| Epi 6h | 5332.74 | 1145.67 | 0.66969 | 0.12292 |
| SA 6h | 10270.3 | 1123.46 | 1.53637 | 0.300961 |
| Me-JA 6h | 9999.32 | 704.628 | 1.49795 | 0.0864855 |
Source Transcript PGSC0003DMT400064455 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27520.1 | +2 | 2e-19 | 86 | 40/72 (56%) | early nodulin-like protein 2 | chr4:13750668-13751819 REVERSE LENGTH=349 |
