Probe CUST_43307_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43307_PI426222305 | JHI_St_60k_v1 | DMT400064364 | ATGGAAGCTTCAACCAGAATAGCGACCTCAAGTTATTAAGAACTGAAAGAAATGCAATGA |
All Microarray Probes Designed to Gene DMG400025005
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43307_PI426222305 | JHI_St_60k_v1 | DMT400064364 | ATGGAAGCTTCAACCAGAATAGCGACCTCAAGTTATTAAGAACTGAAAGAAATGCAATGA |
Microarray Signals from CUST_43307_PI426222305
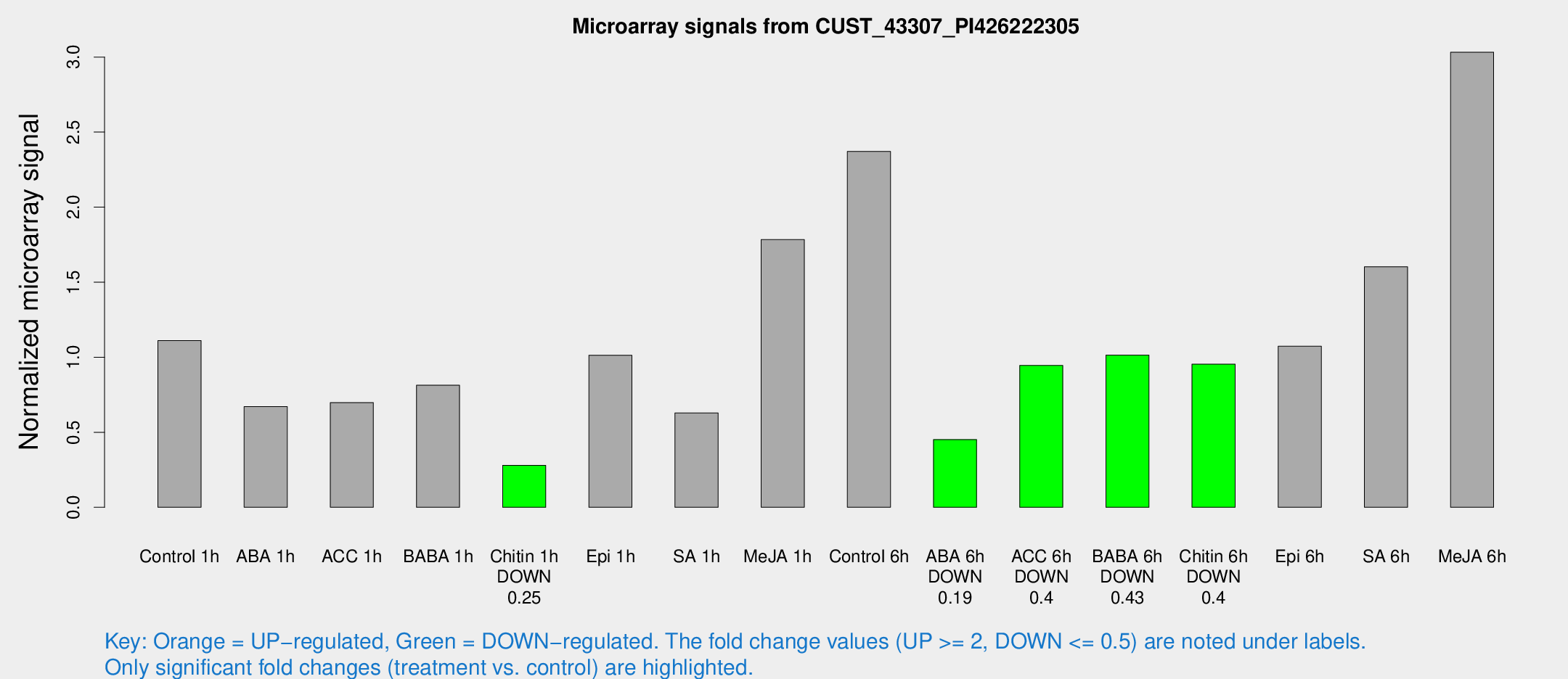
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 32.1727 | 6.49749 | 1.11122 | 0.176959 |
| ABA 1h | 17.0683 | 3.88111 | 0.671072 | 0.16233 |
| ACC 1h | 24.4901 | 8.62854 | 0.697366 | 0.344253 |
| BABA 1h | 22.2742 | 4.1162 | 0.813095 | 0.155062 |
| Chitin 1h | 7.01346 | 3.73911 | 0.27968 | 0.149472 |
| Epi 1h | 25.9674 | 6.74176 | 1.01301 | 0.263575 |
| SA 1h | 19.1453 | 4.70215 | 0.628298 | 0.194395 |
| Me-JA 1h | 41.258 | 5.55566 | 1.78383 | 0.195055 |
| Control 6h | 67.9915 | 10.658 | 2.37139 | 0.203299 |
| ABA 6h | 14.2899 | 4.13466 | 0.451544 | 0.149619 |
| ACC 6h | 30.2108 | 4.96221 | 0.944976 | 0.155227 |
| BABA 6h | 32.0491 | 4.59554 | 1.01375 | 0.147993 |
| Chitin 6h | 28.8075 | 4.66093 | 0.95362 | 0.161035 |
| Epi 6h | 38.4592 | 11.6277 | 1.07308 | 0.497469 |
| SA 6h | 44.2331 | 4.78145 | 1.60303 | 0.17391 |
| Me-JA 6h | 86.6848 | 15.6232 | 3.03204 | 0.40444 |
Source Transcript PGSC0003DMT400064364 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G55290.1 | +3 | 2e-67 | 211 | 106/160 (66%) | NAD(P)-binding Rossmann-fold superfamily protein | chr3:20502653-20503730 FORWARD LENGTH=280 |
