Probe CUST_43225_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43225_PI426222305 | JHI_St_60k_v1 | DMT400002088 | CAAACATCTAATTCAGGATTGGAATTGGACTTGAGATTAGGGCTATAATGTATAGTAGGA |
All Microarray Probes Designed to Gene DMG401000796
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43242_PI426222305 | JHI_St_60k_v1 | DMT400002087 | CAAACATCTAATTCAGGATTGGAATTGGACTTGAGATTAGGGCTATAATGTATAGTAGGA |
| CUST_43225_PI426222305 | JHI_St_60k_v1 | DMT400002088 | CAAACATCTAATTCAGGATTGGAATTGGACTTGAGATTAGGGCTATAATGTATAGTAGGA |
Microarray Signals from CUST_43225_PI426222305
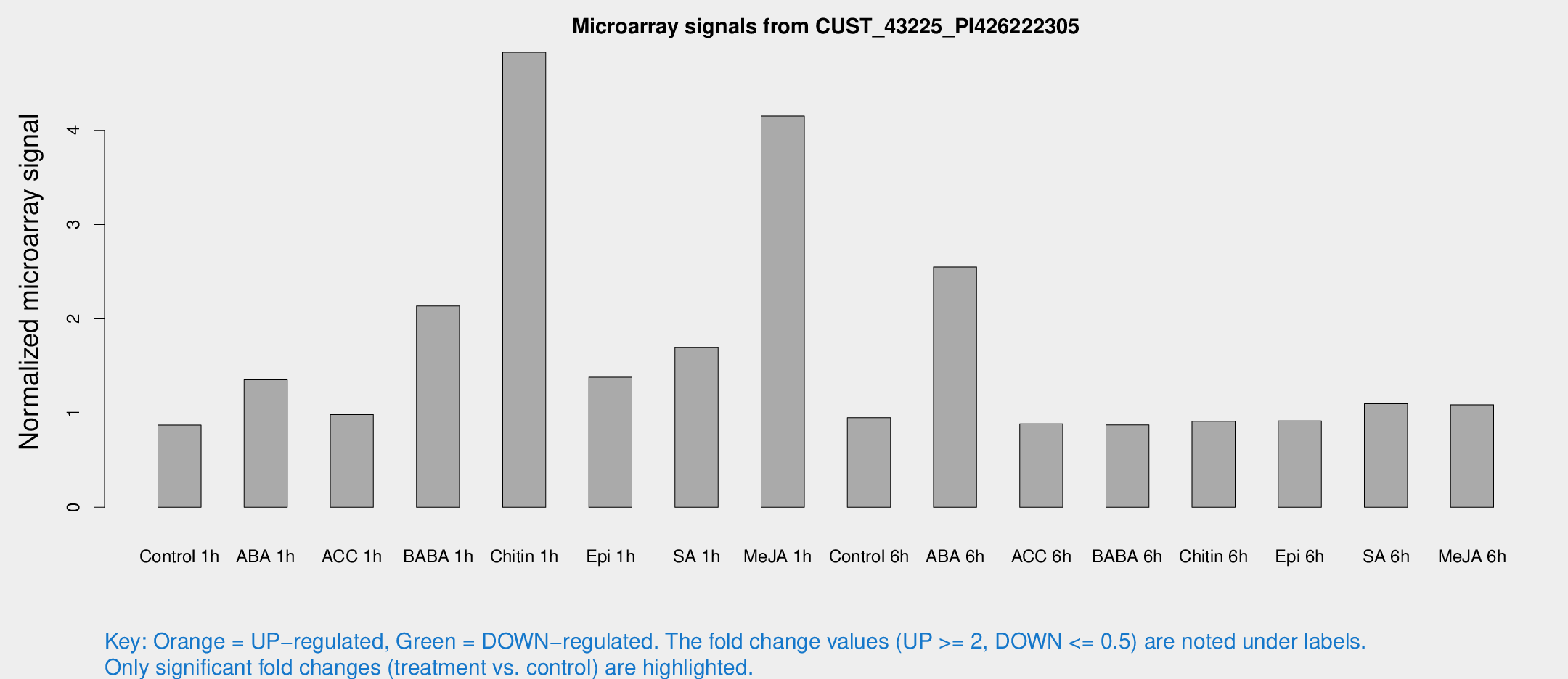
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.17658 | 2.96278 | 0.872538 | 0.498999 |
| ABA 1h | 7.96688 | 3.1891 | 1.35382 | 0.61075 |
| ACC 1h | 6.01339 | 3.2741 | 0.98431 | 0.537324 |
| BABA 1h | 14.0617 | 4.34282 | 2.13584 | 0.719902 |
| Chitin 1h | 26.1648 | 3.32588 | 4.82908 | 0.621095 |
| Epi 1h | 7.75723 | 2.97159 | 1.38131 | 0.635249 |
| SA 1h | 11.5797 | 3.37979 | 1.69352 | 0.596181 |
| Me-JA 1h | 20.1447 | 3.2166 | 4.15201 | 0.664444 |
| Control 6h | 5.67957 | 3.03061 | 0.951345 | 0.512833 |
| ABA 6h | 20.2578 | 9.60862 | 2.55135 | 1.24604 |
| ACC 6h | 6.16874 | 3.66161 | 0.885588 | 0.512906 |
| BABA 6h | 5.81209 | 3.37163 | 0.873825 | 0.506064 |
| Chitin 6h | 5.76062 | 3.34051 | 0.911912 | 0.528653 |
| Epi 6h | 6.17814 | 3.49839 | 0.916376 | 0.514759 |
| SA 6h | 6.58537 | 3.2006 | 1.09868 | 0.559761 |
| Me-JA 6h | 6.78706 | 3.01545 | 1.08827 | 0.535619 |
Source Transcript PGSC0003DMT400002088 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G38470.1 | +1 | 1e-107 | 350 | 178/318 (56%) | ACT-like protein tyrosine kinase family protein | chr4:17999432-18003551 FORWARD LENGTH=575 |
