Probe CUST_43078_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43078_PI426222305 | JHI_St_60k_v1 | DMT400045725 | GTACTGTACCTGATATATGTGCATGTGCAGGCAAAGTATAAGGCAAGAACAATTATAATT |
All Microarray Probes Designed to Gene DMG400017732
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43078_PI426222305 | JHI_St_60k_v1 | DMT400045725 | GTACTGTACCTGATATATGTGCATGTGCAGGCAAAGTATAAGGCAAGAACAATTATAATT |
| CUST_43097_PI426222305 | JHI_St_60k_v1 | DMT400045724 | CATGGTCCACAAATTTTGCTAGAAAGATCTGATTCTTCAAGTCTTATGAGACATAGGTTA |
Microarray Signals from CUST_43078_PI426222305
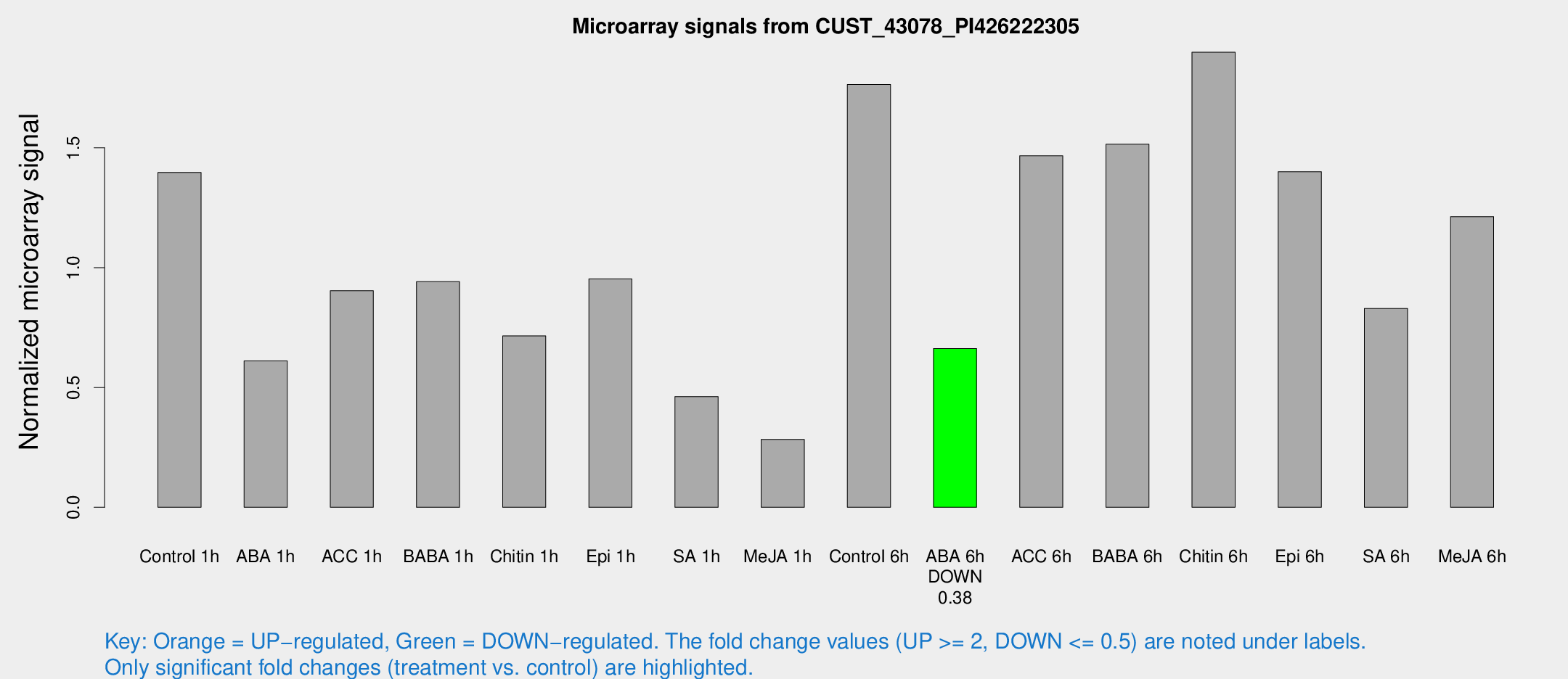
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 177.264 | 32.9371 | 1.39718 | 0.220643 |
| ABA 1h | 67.5674 | 11.1456 | 0.610801 | 0.0949632 |
| ACC 1h | 114.809 | 14.5645 | 0.903659 | 0.0639772 |
| BABA 1h | 115.04 | 21.4061 | 0.941463 | 0.0997128 |
| Chitin 1h | 80.1398 | 12.0087 | 0.715316 | 0.0938856 |
| Epi 1h | 101.115 | 8.08596 | 0.953227 | 0.0756101 |
| SA 1h | 59.1546 | 8.97747 | 0.462302 | 0.09242 |
| Me-JA 1h | 30.4083 | 7.47265 | 0.283433 | 0.0624845 |
| Control 6h | 236.592 | 65.6218 | 1.76441 | 0.437016 |
| ABA 6h | 86.6596 | 8.88145 | 0.662714 | 0.0494858 |
| ACC 6h | 211.994 | 37.5486 | 1.46674 | 0.150004 |
| BABA 6h | 216.035 | 46.7565 | 1.51522 | 0.298181 |
| Chitin 6h | 249.17 | 28.042 | 1.89922 | 0.165704 |
| Epi 6h | 195.017 | 23.8347 | 1.40033 | 0.164455 |
| SA 6h | 113.48 | 39.3458 | 0.829516 | 0.230055 |
| Me-JA 6h | 166.648 | 51.3246 | 1.21306 | 0.379949 |
Source Transcript PGSC0003DMT400045725 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G05300.1 | +1 | 2e-34 | 127 | 63/101 (62%) | zinc transporter 5 precursor | chr1:1545258-1547709 REVERSE LENGTH=360 |
