Probe CUST_43022_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43022_PI426222305 | JHI_St_60k_v1 | DMT400018860 | GAATGTGTACAAAAAAGAATCATTTTTCCTACAACCAAGTTAGACGCACTTAGAGCTAAG |
All Microarray Probes Designed to Gene DMG400007312
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_43069_PI426222305 | JHI_St_60k_v1 | DMT400018861 | GGCTTTGTTTCCGTTCATCTAATTAATATTTACTAGTTCATCTTGGGGTGGTTCAATGAA |
| CUST_43022_PI426222305 | JHI_St_60k_v1 | DMT400018860 | GAATGTGTACAAAAAAGAATCATTTTTCCTACAACCAAGTTAGACGCACTTAGAGCTAAG |
Microarray Signals from CUST_43022_PI426222305
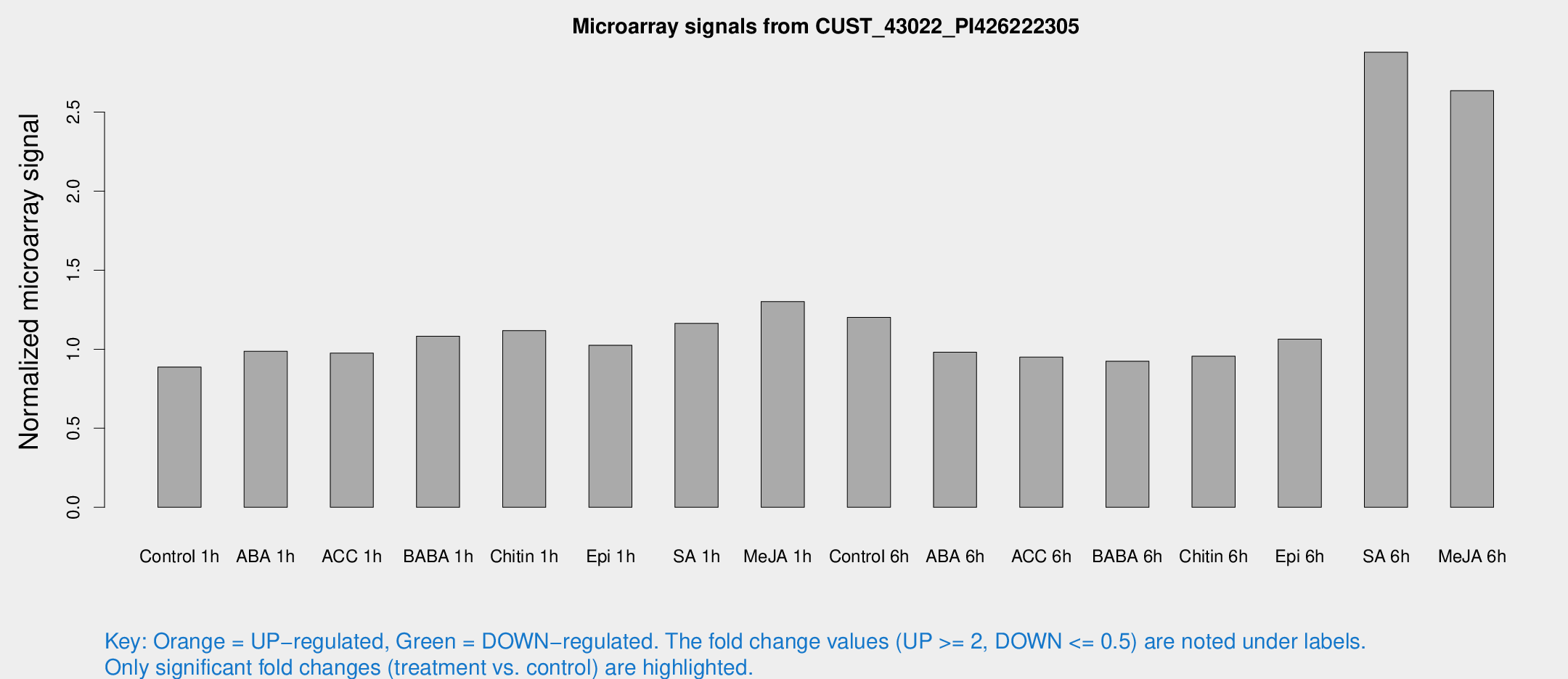
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.35051 | 3.67756 | 0.887561 | 0.513945 |
| ABA 1h | 6.26151 | 3.63278 | 0.98746 | 0.571831 |
| ACC 1h | 7.20177 | 4.19315 | 0.975084 | 0.564855 |
| BABA 1h | 7.51036 | 3.97815 | 1.08255 | 0.581082 |
| Chitin 1h | 7.20385 | 3.81411 | 1.11791 | 0.595188 |
| Epi 1h | 6.35352 | 3.68518 | 1.02496 | 0.593908 |
| SA 1h | 8.73731 | 3.71497 | 1.16316 | 0.518048 |
| Me-JA 1h | 7.76043 | 3.77036 | 1.30178 | 0.660884 |
| Control 6h | 9.16078 | 3.94022 | 1.20142 | 0.58505 |
| ABA 6h | 7.47406 | 4.05381 | 0.980745 | 0.53349 |
| ACC 6h | 8.01087 | 4.7738 | 0.950103 | 0.550639 |
| BABA 6h | 7.4329 | 4.32669 | 0.924174 | 0.535597 |
| Chitin 6h | 7.2811 | 4.21958 | 0.955756 | 0.553717 |
| Epi 6h | 8.72855 | 4.57987 | 1.06361 | 0.565307 |
| SA 6h | 61.9257 | 54.9064 | 2.87865 | 7.49246 |
| Me-JA 6h | 20.8257 | 5.7344 | 2.63501 | 1.05246 |
Source Transcript PGSC0003DMT400018860 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26040.1 | +1 | 1e-50 | 177 | 121/352 (34%) | HXXXD-type acyl-transferase family protein | chr3:9519741-9521069 FORWARD LENGTH=442 |
