Probe CUST_42728_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42728_PI426222305 | JHI_St_60k_v1 | DMT400052362 | AAGTCGTATCCAGCTTCTTCTGTTAATGCACATTCCCCCGCTTCATCTGTAGCAGAATCA |
All Microarray Probes Designed to Gene DMG400020326
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42728_PI426222305 | JHI_St_60k_v1 | DMT400052362 | AAGTCGTATCCAGCTTCTTCTGTTAATGCACATTCCCCCGCTTCATCTGTAGCAGAATCA |
| CUST_42739_PI426222305 | JHI_St_60k_v1 | DMT400052363 | GACATGTCTTAAGATATCAGCCATGGTATCCTGATCTTATTGTTTACTTCCAATGCTAAA |
Microarray Signals from CUST_42728_PI426222305
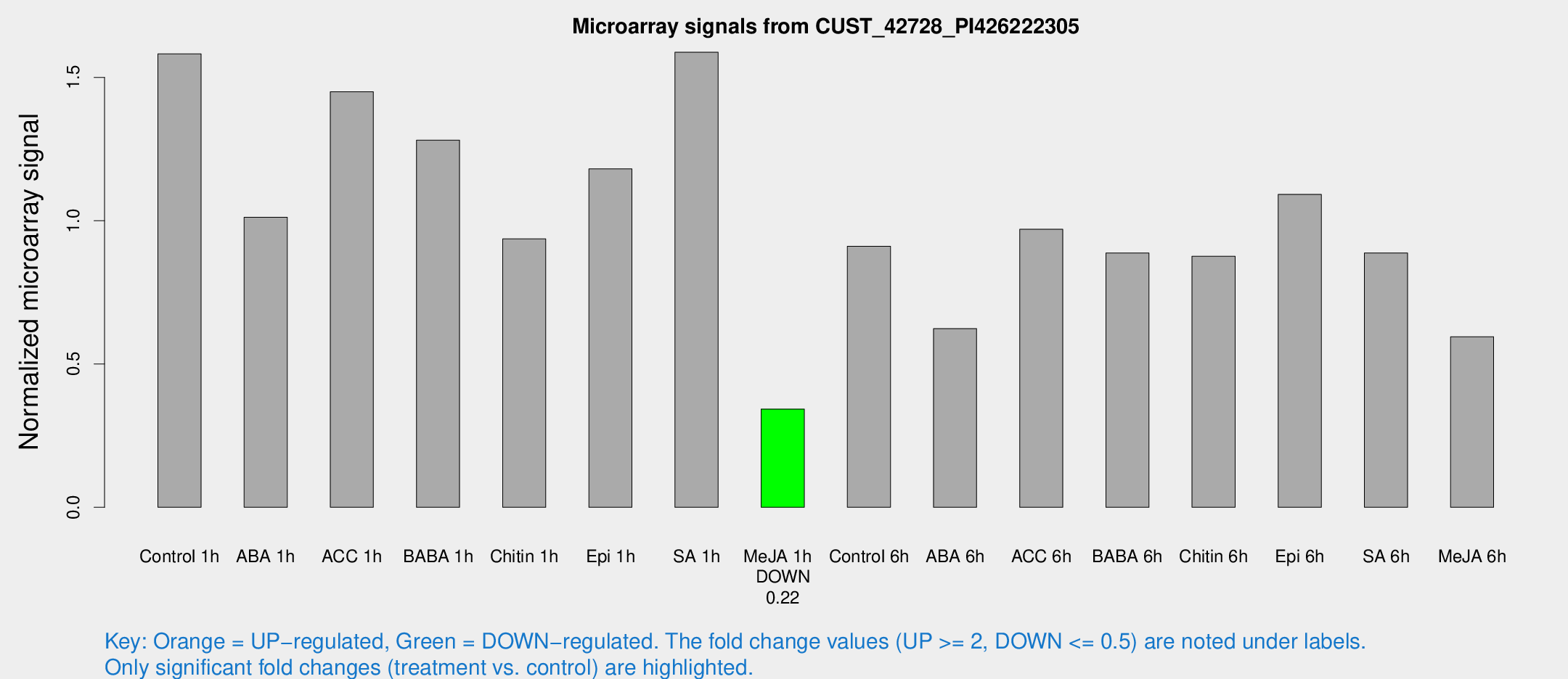
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1898.96 | 294.638 | 1.58189 | 0.136333 |
| ABA 1h | 1053.43 | 61.0982 | 1.01209 | 0.0940088 |
| ACC 1h | 1840.22 | 382.646 | 1.44974 | 0.235725 |
| BABA 1h | 1483.62 | 225.589 | 1.28113 | 0.0939253 |
| Chitin 1h | 1019.19 | 179.02 | 0.936726 | 0.147832 |
| Epi 1h | 1207.75 | 105.794 | 1.1807 | 0.104343 |
| SA 1h | 1932.88 | 200.923 | 1.58788 | 0.091714 |
| Me-JA 1h | 329.131 | 19.3243 | 0.342859 | 0.0288493 |
| Control 6h | 1131.16 | 241.659 | 0.91094 | 0.136829 |
| ABA 6h | 783.023 | 66.2825 | 0.623456 | 0.0361008 |
| ACC 6h | 1335.33 | 200.734 | 0.970062 | 0.0940116 |
| BABA 6h | 1174.18 | 103.909 | 0.887244 | 0.0513001 |
| Chitin 6h | 1097.88 | 70.9949 | 0.875835 | 0.0658605 |
| Epi 6h | 1464.98 | 172.526 | 1.09183 | 0.196489 |
| SA 6h | 1105.36 | 292.397 | 0.887519 | 0.159167 |
| Me-JA 6h | 766.18 | 212.166 | 0.595134 | 0.153733 |
Source Transcript PGSC0003DMT400052362 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37710.1 | +1 | 0.0 | 532 | 283/437 (65%) | receptor lectin kinase | chr2:15814934-15816961 REVERSE LENGTH=675 |
