Probe CUST_42717_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42717_PI426222305 | JHI_St_60k_v1 | DMT400052347 | CTCCATCATGGCTTATAAATTCATTAAGAGATTGTAGGAAGGACCACAAAGTACATCTTT |
All Microarray Probes Designed to Gene DMG400020321
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42717_PI426222305 | JHI_St_60k_v1 | DMT400052347 | CTCCATCATGGCTTATAAATTCATTAAGAGATTGTAGGAAGGACCACAAAGTACATCTTT |
Microarray Signals from CUST_42717_PI426222305
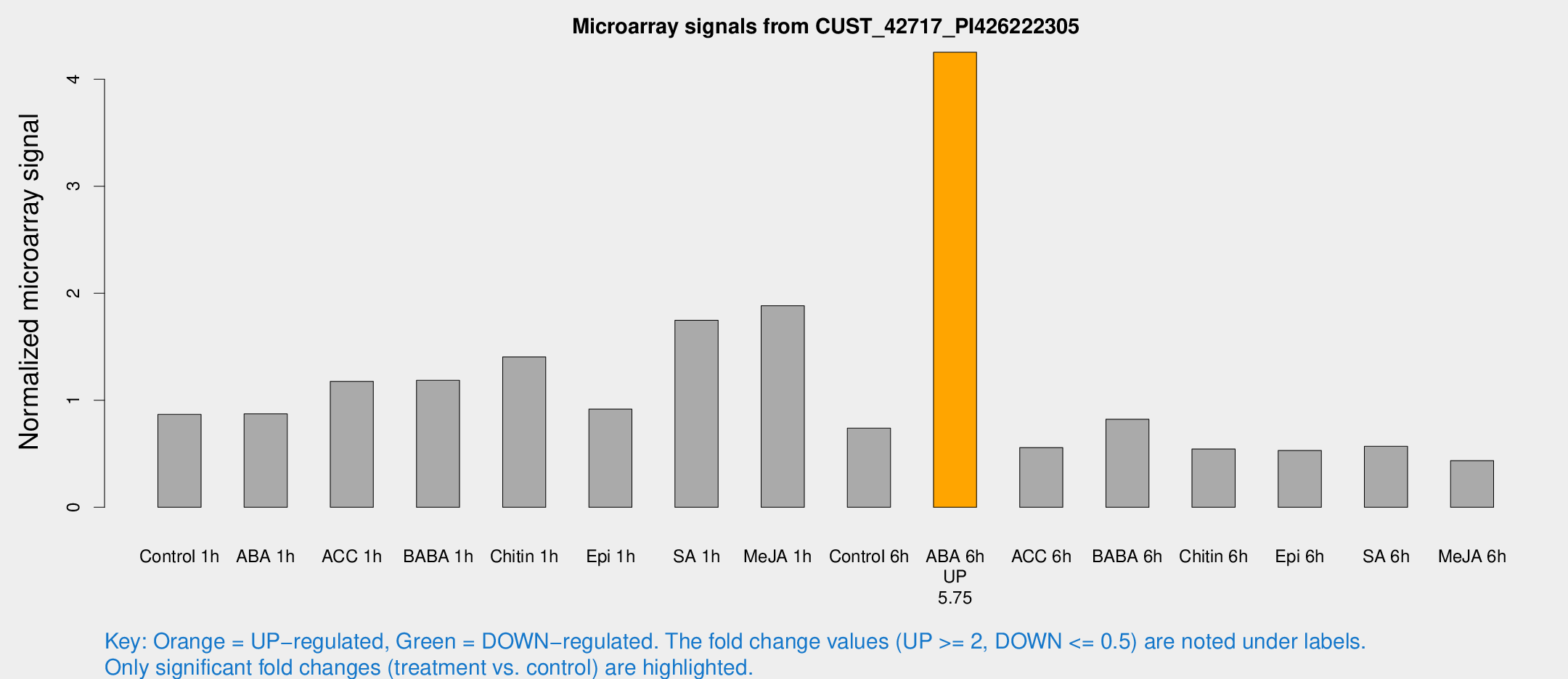
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 20.7681 | 4.73239 | 0.867754 | 0.177972 |
| ABA 1h | 22.6737 | 8.56386 | 0.873163 | 0.656263 |
| ACC 1h | 37.8758 | 16.0934 | 1.17654 | 0.860031 |
| BABA 1h | 28.7049 | 9.01142 | 1.18646 | 0.55 |
| Chitin 1h | 29.1064 | 4.08851 | 1.40535 | 0.200572 |
| Epi 1h | 19.3057 | 5.10865 | 0.917891 | 0.253179 |
| SA 1h | 50.2316 | 22.1589 | 1.74816 | 1.02018 |
| Me-JA 1h | 35.1167 | 4.27889 | 1.88336 | 0.229936 |
| Control 6h | 20.1718 | 6.80618 | 0.739091 | 0.302894 |
| ABA 6h | 104.53 | 11.9438 | 4.25265 | 0.45149 |
| ACC 6h | 21.3251 | 13.004 | 0.558 | 0.5489 |
| BABA 6h | 26.0725 | 9.4071 | 0.822641 | 0.543764 |
| Chitin 6h | 13.4101 | 4.27128 | 0.545037 | 0.179838 |
| Epi 6h | 14.4964 | 4.51895 | 0.530428 | 0.197313 |
| SA 6h | 13.732 | 4.07008 | 0.569705 | 0.234585 |
| Me-JA 6h | 11.596 | 4.854 | 0.435557 | 0.193998 |
Source Transcript PGSC0003DMT400052347 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G38060.1 | +2 | 0.0 | 698 | 359/506 (71%) | phosphate transporter 4;2 | chr2:15922727-15925623 REVERSE LENGTH=512 |
