Probe CUST_42437_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42437_PI426222305 | JHI_St_60k_v1 | DMT400052884 | ATGGTATTTTTCGATGTCACGTCATTAAAGAATGACCGCGATATATCACTTTTCGGCTTG |
All Microarray Probes Designed to Gene DMG400020518
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42437_PI426222305 | JHI_St_60k_v1 | DMT400052884 | ATGGTATTTTTCGATGTCACGTCATTAAAGAATGACCGCGATATATCACTTTTCGGCTTG |
| CUST_42387_PI426222305 | JHI_St_60k_v1 | DMT400052883 | ATGGTATTTTTCGATGTCACGTCATTAAAGAATGACCGCGATATATCACTTTTCGGCTTG |
Microarray Signals from CUST_42437_PI426222305
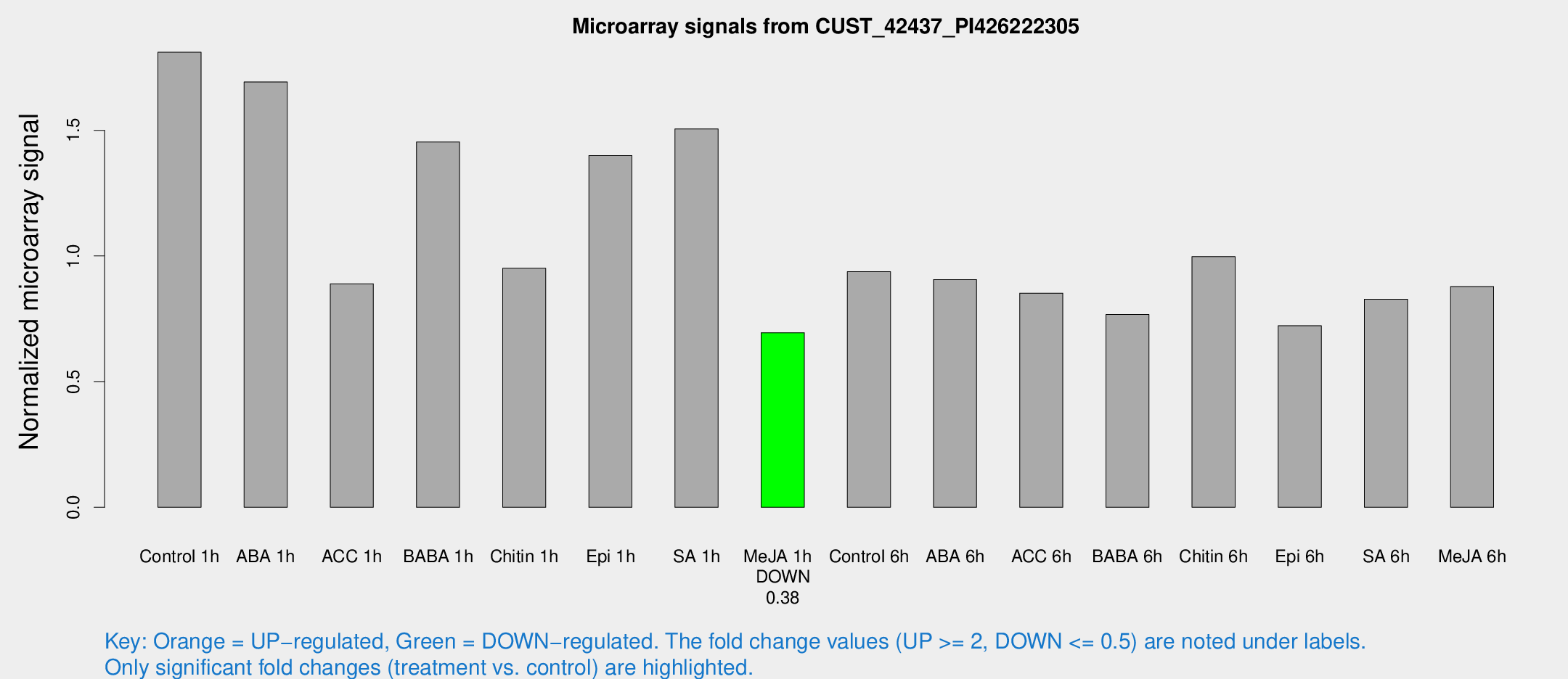
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 754.262 | 43.8233 | 1.81067 | 0.104897 |
| ABA 1h | 671.333 | 174.6 | 1.69257 | 0.380673 |
| ACC 1h | 476.604 | 175.934 | 0.889058 | 0.479509 |
| BABA 1h | 606.908 | 113.84 | 1.4538 | 0.157273 |
| Chitin 1h | 359.225 | 38.4622 | 0.951035 | 0.0557721 |
| Epi 1h | 513.62 | 69.7642 | 1.39964 | 0.191713 |
| SA 1h | 642.972 | 37.328 | 1.50548 | 0.0873242 |
| Me-JA 1h | 238.726 | 28.2823 | 0.694748 | 0.0415945 |
| Control 6h | 407.48 | 82.7987 | 0.937279 | 0.135637 |
| ABA 6h | 417.285 | 78.5368 | 0.906154 | 0.161208 |
| ACC 6h | 408.99 | 35.5779 | 0.852011 | 0.0500558 |
| BABA 6h | 376.006 | 87.6985 | 0.766992 | 0.164969 |
| Chitin 6h | 443.056 | 30.8319 | 0.997348 | 0.0584254 |
| Epi 6h | 345.304 | 50.3755 | 0.72278 | 0.115205 |
| SA 6h | 355.132 | 73.1831 | 0.828076 | 0.100357 |
| Me-JA 6h | 377.114 | 67.1593 | 0.878132 | 0.112079 |
Source Transcript PGSC0003DMT400052884 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G17970.1 | +3 | 0.0 | 663 | 352/522 (67%) | aluminum-activated, malate transporter 12 | chr4:9975482-9977722 FORWARD LENGTH=560 |
