Probe CUST_42364_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42364_PI426222305 | JHI_St_60k_v1 | DMT400011564 | TGAATCAAGTTTGTGTTAATTGTTGCACTGCAGGAGAGGGTTGCAAACTCTATGGATATG |
All Microarray Probes Designed to Gene DMG400004548
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42332_PI426222305 | JHI_St_60k_v1 | DMT400011563 | TATCTACAACATAAATGGCTGTTCACAAAGAAGTTAGTTCCCTTGCTTACCTACTTGTTC |
| CUST_42364_PI426222305 | JHI_St_60k_v1 | DMT400011564 | TGAATCAAGTTTGTGTTAATTGTTGCACTGCAGGAGAGGGTTGCAAACTCTATGGATATG |
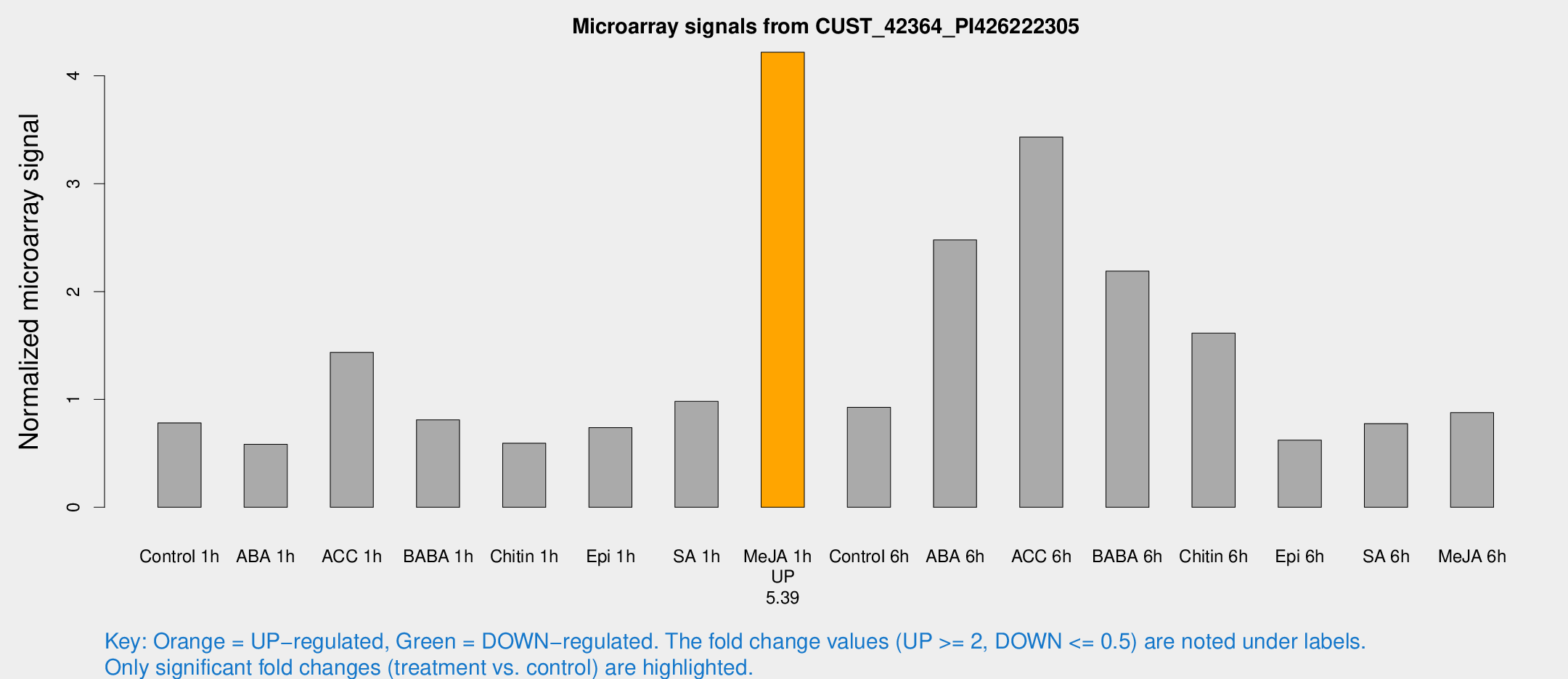
Microarray Signals from CUST_42364_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.44295 | 3.63834 | 0.782392 | 0.328114 |
| ABA 1h | 5.99398 | 3.48477 | 0.584279 | 0.338519 |
| ACC 1h | 19.829 | 8.08836 | 1.4356 | 0.508844 |
| BABA 1h | 10.0914 | 3.80332 | 0.810154 | 0.382445 |
| Chitin 1h | 6.1885 | 3.59473 | 0.594286 | 0.344471 |
| Epi 1h | 7.62855 | 3.51826 | 0.739119 | 0.366695 |
| SA 1h | 11.773 | 3.65095 | 0.98286 | 0.310338 |
| Me-JA 1h | 40.5242 | 5.56538 | 4.21854 | 0.465989 |
| Control 6h | 12.0723 | 4.18945 | 0.926805 | 0.362224 |
| ABA 6h | 31.2686 | 5.13396 | 2.47882 | 0.352957 |
| ACC 6h | 55.5306 | 20.9835 | 3.43228 | 2.02813 |
| BABA 6h | 37.2181 | 19.4464 | 2.18904 | 1.25995 |
| Chitin 6h | 22.776 | 8.69809 | 1.61403 | 0.634858 |
| Epi 6h | 8.12905 | 4.37911 | 0.621772 | 0.334147 |
| SA 6h | 9.56851 | 3.88504 | 0.775903 | 0.370582 |
| Me-JA 6h | 11.2235 | 3.76754 | 0.877408 | 0.35815 |
Source Transcript PGSC0003DMT400011564 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G72060.1 | +3 | 1e-05 | 42 | 22/40 (55%) | serine-type endopeptidase inhibitors | chr1:27118494-27118843 FORWARD LENGTH=81 |
