Probe CUST_42292_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42292_PI426222305 | JHI_St_60k_v1 | DMT400016038 | AAAGATGACACTAGCAAAGAGCTGCTTGAATGGGGTTCTAGGGTCTCAGCAGCTAATAGT |
All Microarray Probes Designed to Gene DMG400006270
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42292_PI426222305 | JHI_St_60k_v1 | DMT400016038 | AAAGATGACACTAGCAAAGAGCTGCTTGAATGGGGTTCTAGGGTCTCAGCAGCTAATAGT |
Microarray Signals from CUST_42292_PI426222305
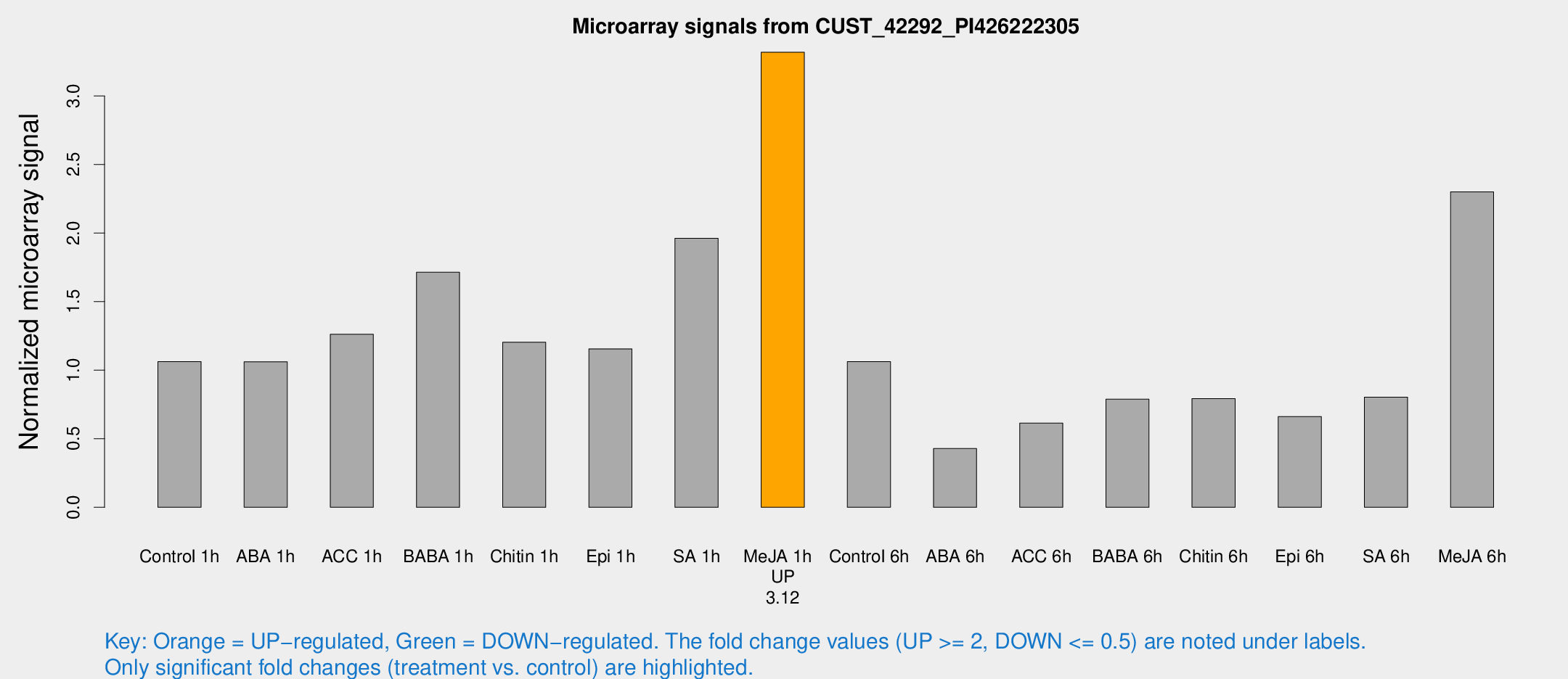
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11253.7 | 1363.19 | 1.06279 | 0.120513 |
| ABA 1h | 10587.2 | 2938.89 | 1.06121 | 0.231033 |
| ACC 1h | 15365.9 | 5679.02 | 1.26149 | 0.416873 |
| BABA 1h | 17892.7 | 3527.09 | 1.71435 | 0.226221 |
| Chitin 1h | 11334.4 | 704.188 | 1.20404 | 0.069516 |
| Epi 1h | 11018.8 | 2327.89 | 1.15556 | 0.278626 |
| SA 1h | 23138.7 | 7504.79 | 1.96199 | 0.665492 |
| Me-JA 1h | 29096.5 | 5085.3 | 3.3187 | 0.28404 |
| Control 6h | 12385.7 | 4279.92 | 1.06213 | 0.286795 |
| ABA 6h | 4828.96 | 601.127 | 0.428544 | 0.0682779 |
| ACC 6h | 7519.17 | 1134.29 | 0.613565 | 0.133995 |
| BABA 6h | 9230.94 | 534.535 | 0.788203 | 0.0584996 |
| Chitin 6h | 9132.02 | 1832.26 | 0.792723 | 0.157392 |
| Epi 6h | 8376.7 | 2042.96 | 0.661865 | 0.2471 |
| SA 6h | 8574.53 | 1459.31 | 0.803482 | 0.0576113 |
| Me-JA 6h | 24271.5 | 3109.21 | 2.30082 | 0.292911 |
Source Transcript PGSC0003DMT400016038 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G33070.1 | +1 | 0.0 | 993 | 503/607 (83%) | Thiamine pyrophosphate dependent pyruvate decarboxylase family protein | chr4:15952519-15954676 REVERSE LENGTH=607 |
