Probe CUST_42078_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42078_PI426222305 | JHI_St_60k_v1 | DMT400050529 | GAAACCAGCACAACCCATACACGCAGCTTTTGGTCCACCTGAAACGAAATCAGATTGTAT |
All Microarray Probes Designed to Gene DMG401019636
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_42078_PI426222305 | JHI_St_60k_v1 | DMT400050529 | GAAACCAGCACAACCCATACACGCAGCTTTTGGTCCACCTGAAACGAAATCAGATTGTAT |
Microarray Signals from CUST_42078_PI426222305
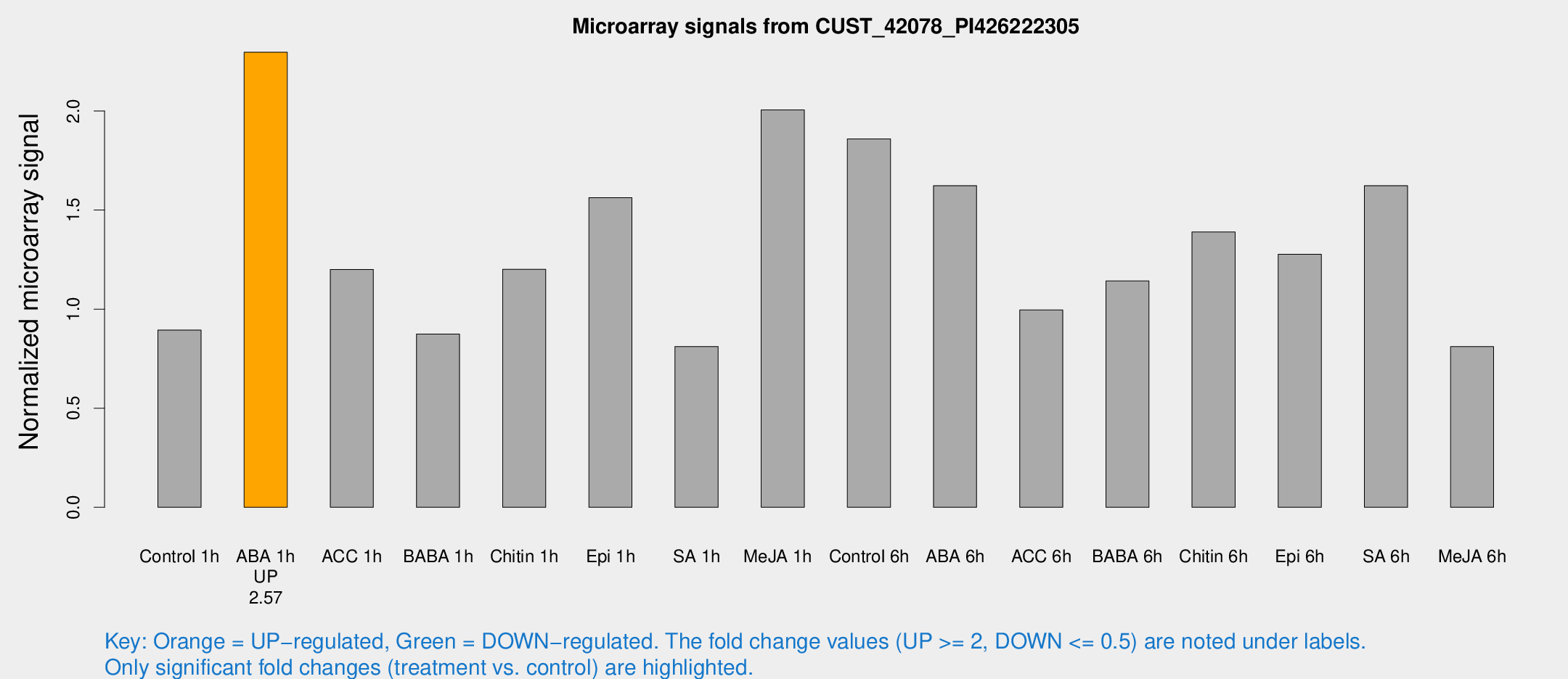
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.89651 | 3.38967 | 0.894166 | 0.461149 |
| ABA 1h | 15.2538 | 3.36931 | 2.29653 | 0.50907 |
| ACC 1h | 9.54274 | 4.33387 | 1.20028 | 0.568584 |
| BABA 1h | 6.31643 | 3.61588 | 0.873936 | 0.499096 |
| Chitin 1h | 9.21465 | 3.46293 | 1.20115 | 0.563749 |
| Epi 1h | 10.8869 | 3.34033 | 1.56272 | 0.546662 |
| SA 1h | 6.24801 | 3.44212 | 0.811203 | 0.447245 |
| Me-JA 1h | 16.2094 | 8.81999 | 2.00572 | 1.56837 |
| Control 6h | 18.9895 | 10.5261 | 1.85943 | 1.11561 |
| ABA 6h | 14.0778 | 3.70498 | 1.6234 | 0.534151 |
| ACC 6h | 8.80639 | 4.16387 | 0.995876 | 0.48159 |
| BABA 6h | 10.7522 | 3.96491 | 1.14232 | 0.535553 |
| Chitin 6h | 12.0246 | 3.94541 | 1.38941 | 0.54657 |
| Epi 6h | 12.1972 | 4.38305 | 1.27737 | 0.591118 |
| SA 6h | 18.2939 | 11.8657 | 1.62318 | 1.94926 |
| Me-JA 6h | 6.0436 | 3.51407 | 0.810807 | 0.471406 |
Source Transcript PGSC0003DMT400050529 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G10110.1 | -1 | 2e-10 | 56 | 24/27 (89%) | Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein | chr3:3116225-3117378 FORWARD LENGTH=173 |
