Probe CUST_41975_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41975_PI426222305 | JHI_St_60k_v1 | DMT400022327 | GAGGGACCTGATGAACCTTTTTTGAATATGCAAATTCTTGACAGGTACGTATACCTAATA |
All Microarray Probes Designed to Gene DMG401008665
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41975_PI426222305 | JHI_St_60k_v1 | DMT400022327 | GAGGGACCTGATGAACCTTTTTTGAATATGCAAATTCTTGACAGGTACGTATACCTAATA |
Microarray Signals from CUST_41975_PI426222305
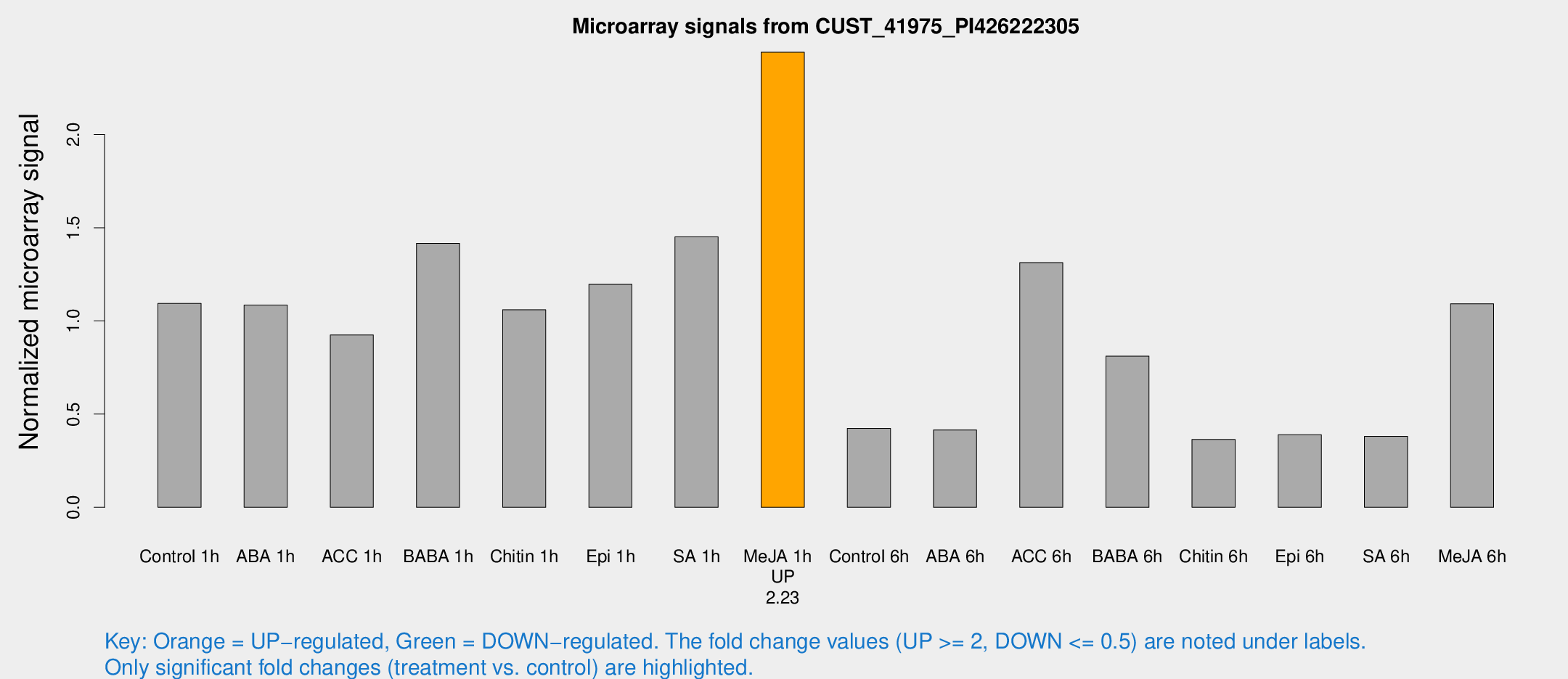
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 725.246 | 84.7802 | 1.09348 | 0.117147 |
| ABA 1h | 633.434 | 58.5536 | 1.08483 | 0.064396 |
| ACC 1h | 687.732 | 214.557 | 0.924794 | 0.256902 |
| BABA 1h | 924.379 | 159.028 | 1.41633 | 0.123197 |
| Chitin 1h | 639.576 | 101.904 | 1.05998 | 0.148056 |
| Epi 1h | 687.067 | 84.4483 | 1.19646 | 0.151179 |
| SA 1h | 1005.83 | 169.198 | 1.45139 | 0.211789 |
| Me-JA 1h | 1328.69 | 184.269 | 2.44136 | 0.141079 |
| Control 6h | 296.106 | 70.1177 | 0.423953 | 0.0783174 |
| ABA 6h | 321.533 | 99.0337 | 0.415095 | 0.125728 |
| ACC 6h | 1003.78 | 140.901 | 1.31231 | 0.299488 |
| BABA 6h | 659.219 | 215.846 | 0.811105 | 0.264609 |
| Chitin 6h | 254.338 | 15.1405 | 0.364124 | 0.0224304 |
| Epi 6h | 307.852 | 79.6311 | 0.38919 | 0.076575 |
| SA 6h | 255.215 | 48.4543 | 0.380289 | 0.0399182 |
| Me-JA 6h | 715.688 | 59.2045 | 1.09168 | 0.181574 |
Source Transcript PGSC0003DMT400022327 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G06570.1 | +1 | 1e-81 | 251 | 125/237 (53%) | alpha/beta-Hydrolases superfamily protein | chr5:2008075-2011013 REVERSE LENGTH=329 |
