Probe CUST_41763_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41763_PI426222305 | JHI_St_60k_v1 | DMT400015593 | CTAGTAGTTGTGGAAAATCCAATTTCAAATAAGAGGATGTTTAAGGTTGGTCAAGTGGAA |
All Microarray Probes Designed to Gene DMG400006088
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41744_PI426222305 | JHI_St_60k_v1 | DMT400015592 | GATCTATTTACTGATGTCTCAGAGGAAGGACTATCTCCAAGGCAAGTCACAGCAAGATAG |
| CUST_41763_PI426222305 | JHI_St_60k_v1 | DMT400015593 | CTAGTAGTTGTGGAAAATCCAATTTCAAATAAGAGGATGTTTAAGGTTGGTCAAGTGGAA |
Microarray Signals from CUST_41763_PI426222305
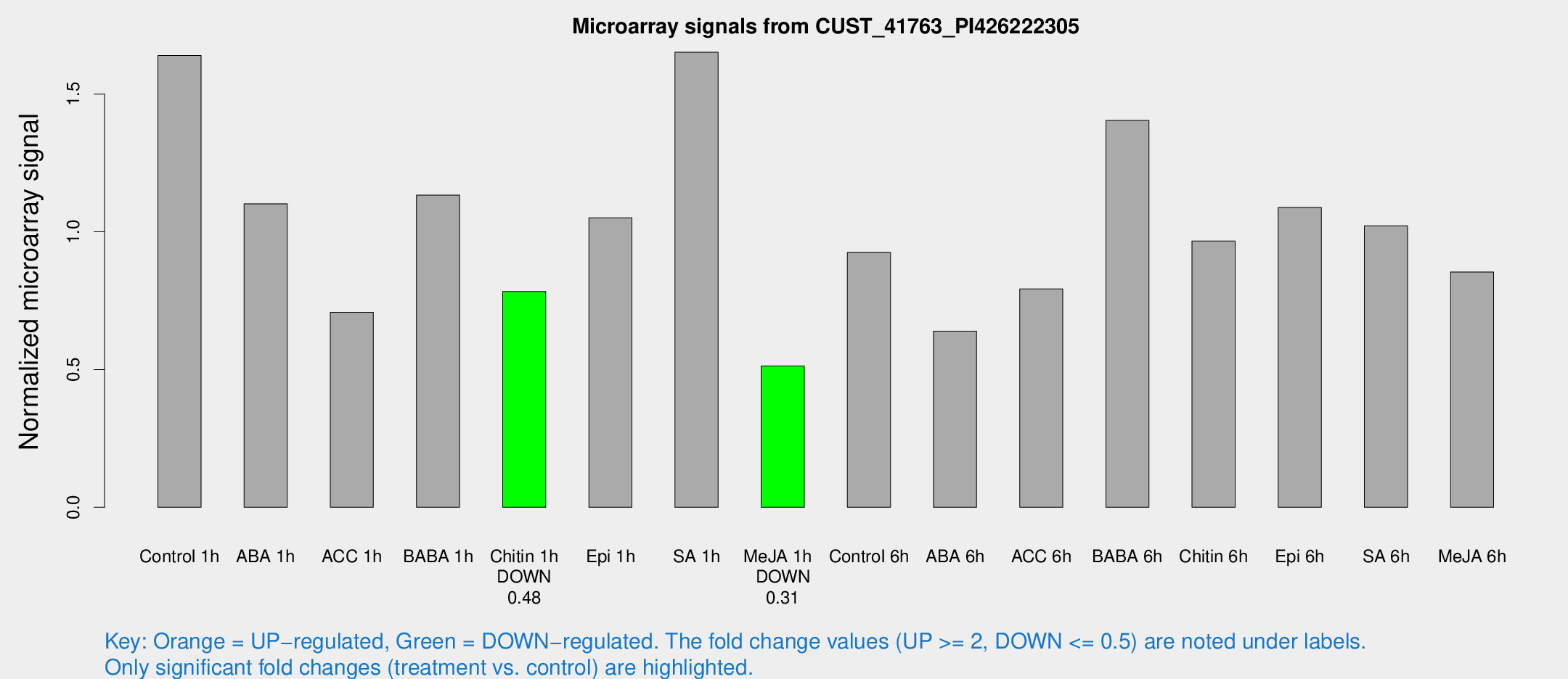
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2212.14 | 341.435 | 1.63975 | 0.16358 |
| ABA 1h | 1324.54 | 211.945 | 1.10168 | 0.140398 |
| ACC 1h | 1088.04 | 325.556 | 0.707726 | 0.233238 |
| BABA 1h | 1520.23 | 317.93 | 1.1332 | 0.155946 |
| Chitin 1h | 953.176 | 154.73 | 0.783497 | 0.0912412 |
| Epi 1h | 1222.91 | 164.87 | 1.05096 | 0.142662 |
| SA 1h | 2266.31 | 256.99 | 1.65181 | 0.114807 |
| Me-JA 1h | 557.029 | 50.7575 | 0.512873 | 0.041346 |
| Control 6h | 1266.2 | 226.229 | 0.925362 | 0.0899665 |
| ABA 6h | 943.792 | 220.289 | 0.639628 | 0.108802 |
| ACC 6h | 1200.62 | 69.4979 | 0.792778 | 0.12009 |
| BABA 6h | 2078.96 | 120.402 | 1.40408 | 0.0811185 |
| Chitin 6h | 1358.53 | 78.6199 | 0.966449 | 0.0558873 |
| Epi 6h | 1629.05 | 129.651 | 1.08816 | 0.0628996 |
| SA 6h | 1467.48 | 405.94 | 1.02169 | 0.222064 |
| Me-JA 6h | 1227.5 | 367.21 | 0.854241 | 0.190696 |
Source Transcript PGSC0003DMT400015593 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G23770.1 | +2 | 5e-79 | 267 | 203/596 (34%) | protein kinase family protein / peptidoglycan-binding LysM domain-containing protein | chr2:10120242-10122080 REVERSE LENGTH=612 |
