Probe CUST_41637_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41637_PI426222305 | JHI_St_60k_v1 | DMT400055477 | CTCCACGACTACACATGGTTAAACTTCTTTCTGAAATCTAATCCTAACCAAGATTCTAAT |
All Microarray Probes Designed to Gene DMG400021541
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41637_PI426222305 | JHI_St_60k_v1 | DMT400055477 | CTCCACGACTACACATGGTTAAACTTCTTTCTGAAATCTAATCCTAACCAAGATTCTAAT |
Microarray Signals from CUST_41637_PI426222305
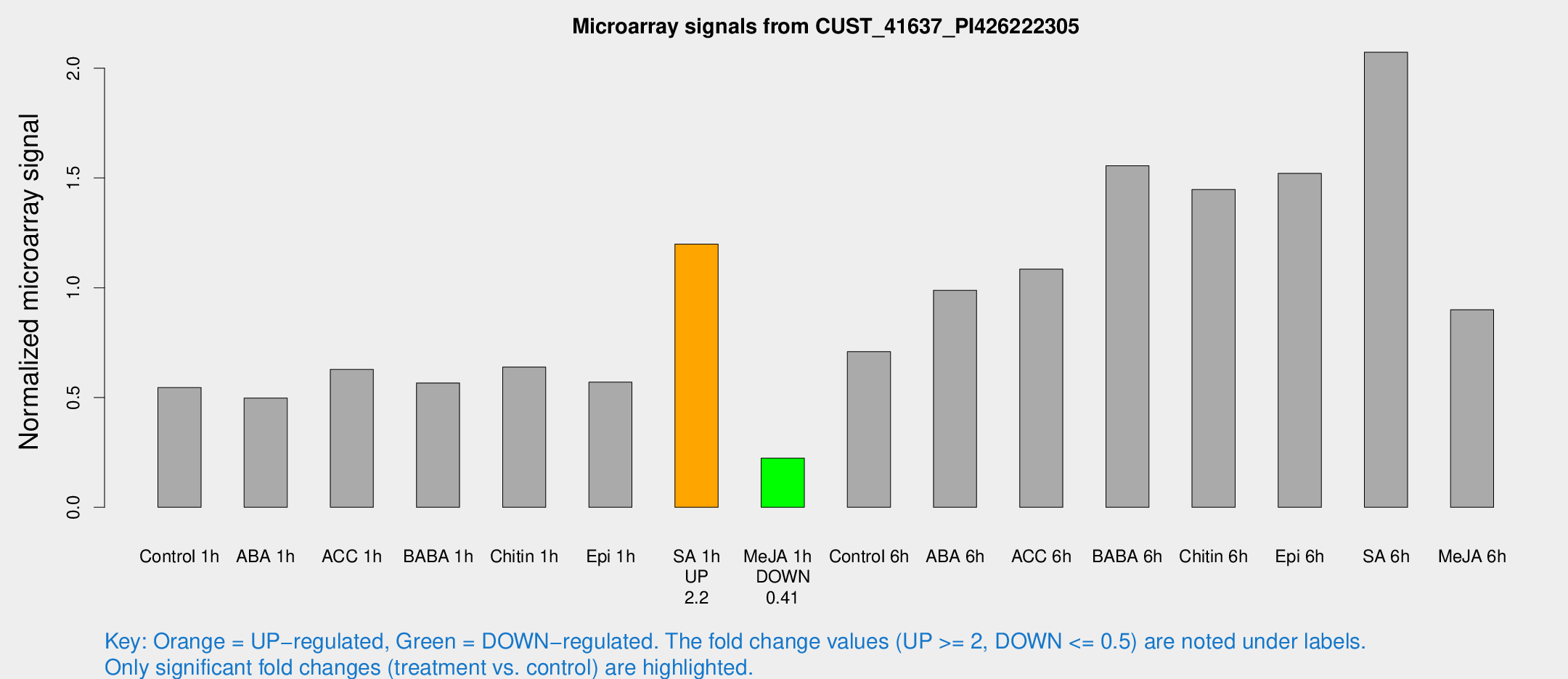
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 20.3893 | 3.71355 | 0.545046 | 0.100499 |
| ABA 1h | 17.225 | 4.20731 | 0.497233 | 0.116802 |
| ACC 1h | 28.1423 | 10.0166 | 0.62789 | 0.253085 |
| BABA 1h | 22.8435 | 6.73521 | 0.566491 | 0.164887 |
| Chitin 1h | 21.3132 | 3.71551 | 0.63854 | 0.111888 |
| Epi 1h | 20.3311 | 6.1213 | 0.569949 | 0.235221 |
| SA 1h | 46.9323 | 8.51393 | 1.19855 | 0.139644 |
| Me-JA 1h | 6.80714 | 3.65071 | 0.223724 | 0.11959 |
| Control 6h | 30.3013 | 9.53429 | 0.709079 | 0.244246 |
| ABA 6h | 42.736 | 12.738 | 0.988486 | 0.305863 |
| ACC 6h | 46.5177 | 5.08024 | 1.08473 | 0.114555 |
| BABA 6h | 66.6186 | 11.5999 | 1.55589 | 0.24919 |
| Chitin 6h | 57.0557 | 5.21443 | 1.44785 | 0.132207 |
| Epi 6h | 64.2606 | 7.75106 | 1.52086 | 0.212656 |
| SA 6h | 79.7758 | 18.7097 | 2.07278 | 0.772277 |
| Me-JA 6h | 36.4093 | 10.2501 | 0.899762 | 0.221816 |
Source Transcript PGSC0003DMT400055477 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37320.1 | +2 | 3e-155 | 458 | 232/478 (49%) | cytochrome P450, family 81, subfamily D, polypeptide 5 | chr4:17559742-17561690 REVERSE LENGTH=495 |
