Probe CUST_41481_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41481_PI426222305 | JHI_St_60k_v1 | DMT400021572 | GTCCCTTTGTGATGAAATTAATACAATGGCTAGTTTGGTTAGAGCTATGGCTATTGAACT |
All Microarray Probes Designed to Gene DMG400008364
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41481_PI426222305 | JHI_St_60k_v1 | DMT400021572 | GTCCCTTTGTGATGAAATTAATACAATGGCTAGTTTGGTTAGAGCTATGGCTATTGAACT |
Microarray Signals from CUST_41481_PI426222305
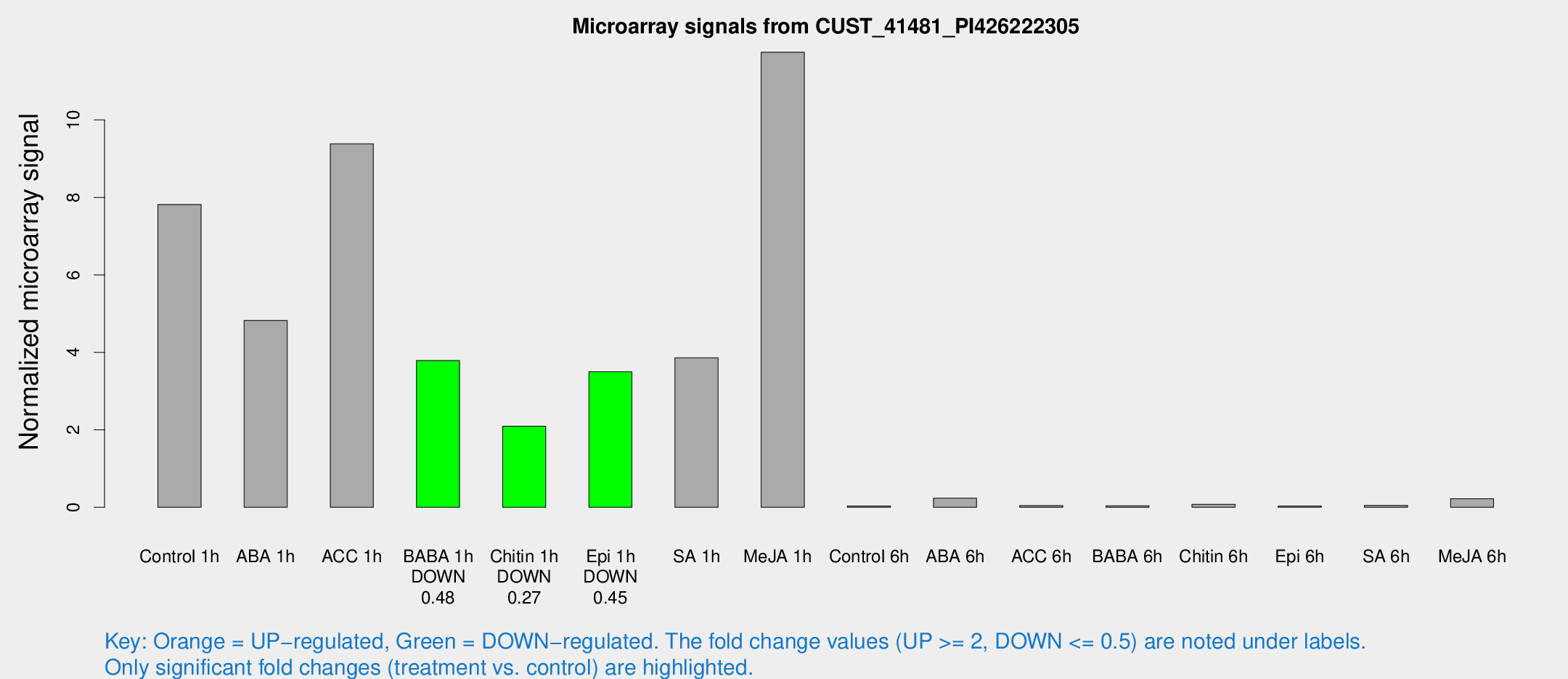
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1492.57 | 308.73 | 7.81606 | 1.13093 |
| ABA 1h | 898.87 | 277.93 | 4.82403 | 1.8836 |
| ACC 1h | 2172.33 | 860.57 | 9.3825 | 4.53856 |
| BABA 1h | 686.159 | 115.56 | 3.78779 | 0.409637 |
| Chitin 1h | 366.299 | 87.5986 | 2.09304 | 0.473 |
| Epi 1h | 557.124 | 44.1966 | 3.49695 | 0.381768 |
| SA 1h | 826.237 | 309.97 | 3.85854 | 1.58867 |
| Me-JA 1h | 1800.53 | 292.642 | 11.7483 | 0.887245 |
| Control 6h | 6.57463 | 3.8203 | 0.0358227 | 0.0207846 |
| ABA 6h | 51.6763 | 17.1136 | 0.23746 | 0.0712767 |
| ACC 6h | 9.88178 | 4.49052 | 0.0457342 | 0.0218898 |
| BABA 6h | 8.60272 | 4.27718 | 0.0402688 | 0.0209762 |
| Chitin 6h | 19.6056 | 10.0269 | 0.0782346 | 0.0478057 |
| Epi 6h | 7.35331 | 4.31932 | 0.0351999 | 0.0203985 |
| SA 6h | 8.94871 | 4.02408 | 0.0487926 | 0.0226811 |
| Me-JA 6h | 45.9924 | 16.2099 | 0.223044 | 0.113502 |
Source Transcript PGSC0003DMT400021572 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G55290.1 | +1 | 7e-41 | 140 | 68/116 (59%) | NAD(P)-binding Rossmann-fold superfamily protein | chr3:20502653-20503730 FORWARD LENGTH=280 |
