Probe CUST_41126_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41126_PI426222305 | JHI_St_60k_v1 | DMT400004379 | GTTTCTGTGTTTGATGACTAATTTGACAGAAAGTTGATCTTAACATCTCCAGTGATGCTT |
All Microarray Probes Designed to Gene DMG400001739
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41126_PI426222305 | JHI_St_60k_v1 | DMT400004379 | GTTTCTGTGTTTGATGACTAATTTGACAGAAAGTTGATCTTAACATCTCCAGTGATGCTT |
Microarray Signals from CUST_41126_PI426222305
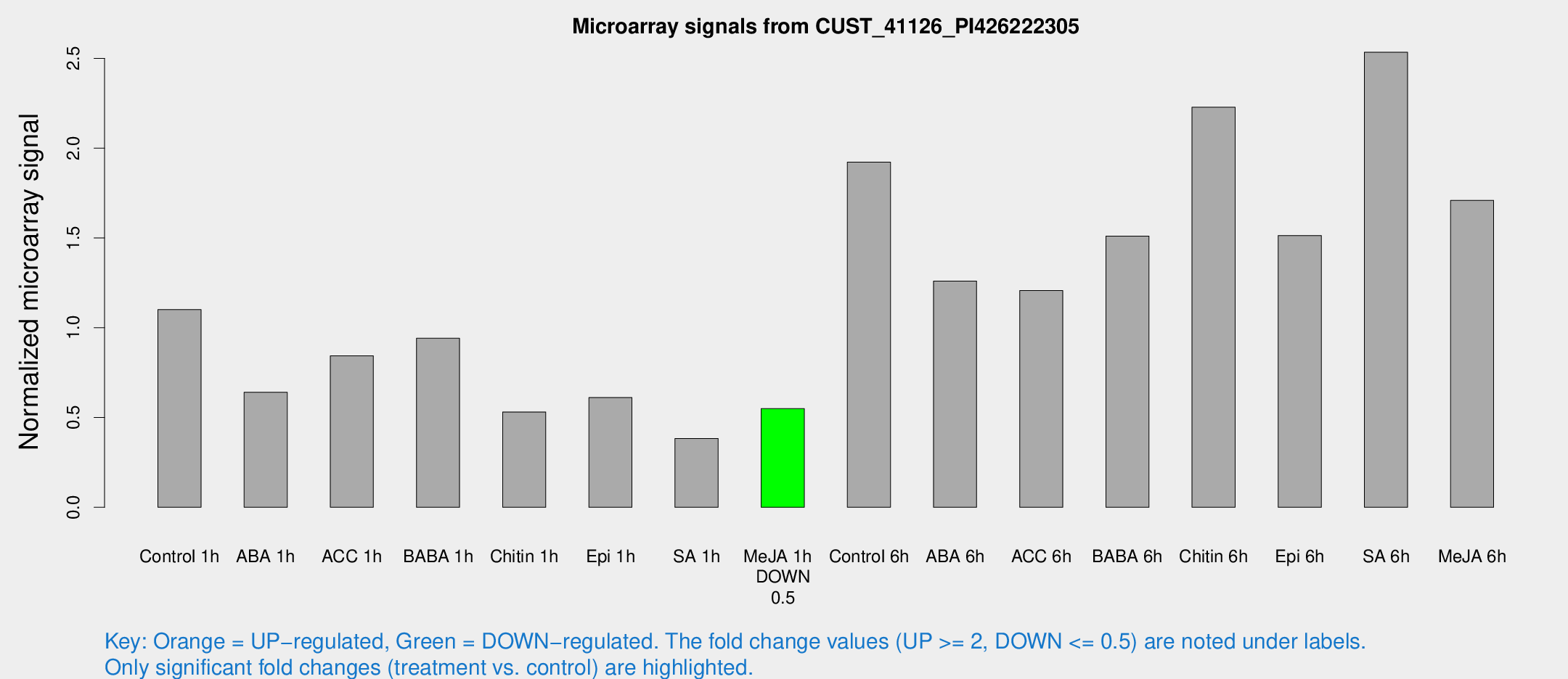
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 20.2373 | 3.21492 | 1.1006 | 0.178604 |
| ABA 1h | 10.5188 | 2.97032 | 0.640626 | 0.189693 |
| ACC 1h | 16.1345 | 3.3882 | 0.843404 | 0.191322 |
| BABA 1h | 16.9021 | 3.33539 | 0.941395 | 0.196826 |
| Chitin 1h | 9.29679 | 3.09488 | 0.530835 | 0.199847 |
| Epi 1h | 9.67917 | 3.08003 | 0.611012 | 0.198297 |
| SA 1h | 7.99922 | 3.055 | 0.38352 | 0.181495 |
| Me-JA 1h | 8.63394 | 3.13251 | 0.549292 | 0.2224 |
| Control 6h | 38.7247 | 10.4965 | 1.92165 | 0.485474 |
| ABA 6h | 27.1157 | 9.07093 | 1.25972 | 0.498889 |
| ACC 6h | 28.5538 | 8.54884 | 1.20664 | 0.4349 |
| BABA 6h | 31.138 | 3.98072 | 1.51036 | 0.194876 |
| Chitin 6h | 43.566 | 4.28858 | 2.22786 | 0.223222 |
| Epi 6h | 35.972 | 11.2954 | 1.5133 | 0.884049 |
| SA 6h | 45.8753 | 4.25213 | 2.53404 | 0.486813 |
| Me-JA 6h | 36.8626 | 15.1323 | 1.70922 | 0.625537 |
Source Transcript PGSC0003DMT400004379 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G09690.1 | +1 | 0.0 | 615 | 289/505 (57%) | alpha/beta-Hydrolases superfamily protein | chr3:2972356-2974592 FORWARD LENGTH=527 |
