Probe CUST_41114_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41114_PI426222305 | JHI_St_60k_v1 | DMT400004447 | CTCGTGGTTGCATTTGTTTAGTTTTCTTGTTTTTTGTCTGTGTTGCATCTTCAAGTGAAA |
All Microarray Probes Designed to Gene DMG400001774
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_41114_PI426222305 | JHI_St_60k_v1 | DMT400004447 | CTCGTGGTTGCATTTGTTTAGTTTTCTTGTTTTTTGTCTGTGTTGCATCTTCAAGTGAAA |
| CUST_41094_PI426222305 | JHI_St_60k_v1 | DMT400004446 | CTCGTGGTTGCATTTGTTTAGTTTTCTTGTTTTTTGTCTGTGTTGCATCTTCAAGTGAAA |
Microarray Signals from CUST_41114_PI426222305
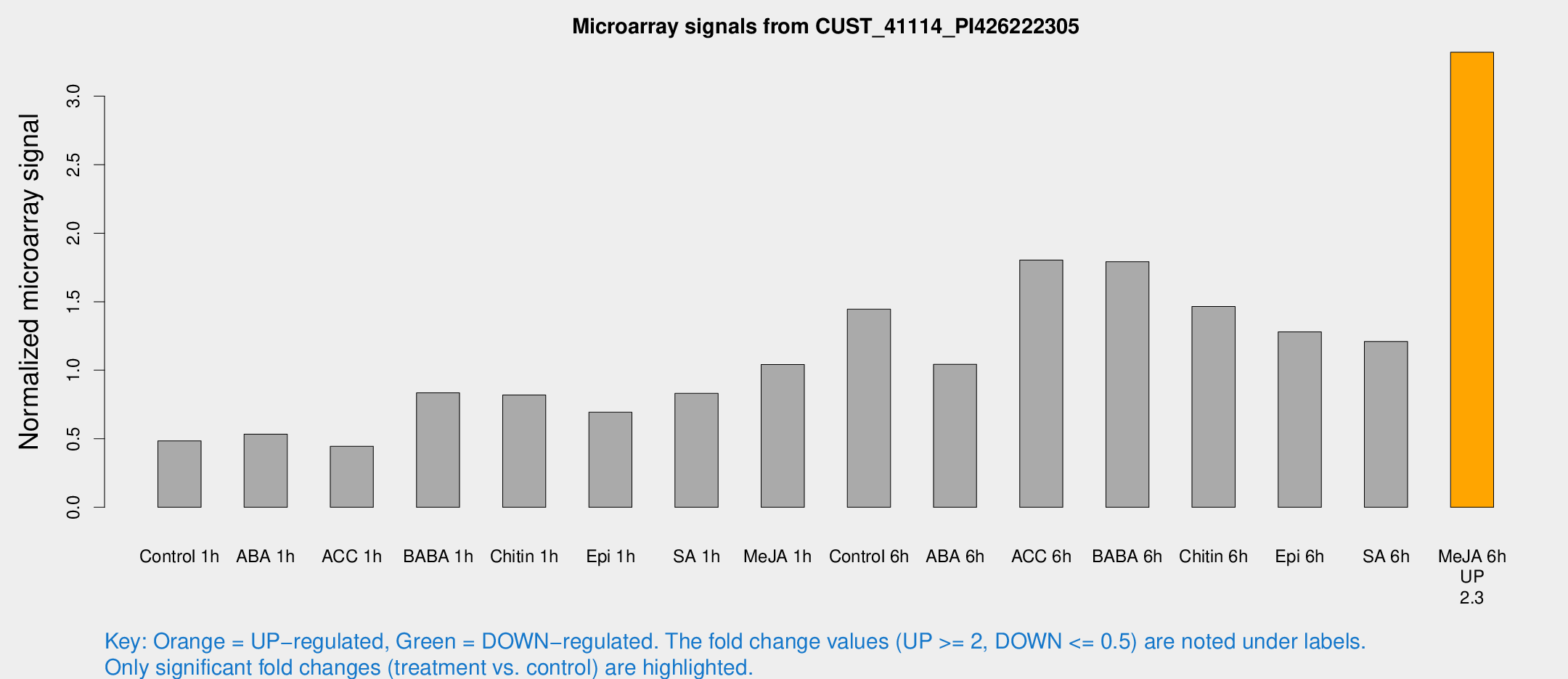
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 457.96 | 26.7153 | 0.484576 | 0.0281925 |
| ABA 1h | 465.008 | 88.4587 | 0.534079 | 0.0895761 |
| ACC 1h | 478.455 | 141.831 | 0.445292 | 0.126033 |
| BABA 1h | 801.841 | 170.927 | 0.83477 | 0.120041 |
| Chitin 1h | 731.555 | 163.407 | 0.819053 | 0.176914 |
| Epi 1h | 582.434 | 98.8271 | 0.693671 | 0.119544 |
| SA 1h | 809.769 | 66.7505 | 0.830792 | 0.0631211 |
| Me-JA 1h | 841.498 | 194.938 | 1.0415 | 0.148776 |
| Control 6h | 1422.11 | 275.304 | 1.44459 | 0.191042 |
| ABA 6h | 1072.35 | 174.238 | 1.04315 | 0.109062 |
| ACC 6h | 2052.51 | 421.523 | 1.80339 | 0.446497 |
| BABA 6h | 1969.26 | 402.827 | 1.79116 | 0.300063 |
| Chitin 6h | 1473.38 | 85.3591 | 1.46504 | 0.0846663 |
| Epi 6h | 1371.11 | 121.023 | 1.27985 | 0.197385 |
| SA 6h | 1186.84 | 258.422 | 1.20989 | 0.155758 |
| Me-JA 6h | 3120.47 | 180.485 | 3.3198 | 0.191703 |
Source Transcript PGSC0003DMT400004447 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G37130.1 | +3 | 6e-162 | 464 | 219/305 (72%) | Peroxidase superfamily protein | chr2:15598225-15600004 REVERSE LENGTH=327 |
