Probe CUST_40937_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40937_PI426222305 | JHI_St_60k_v1 | DMT400079776 | TAATCTCACACTTCCTCATCAACATCCTTACCCTGATGCTGTGGTTCAAGAAGTACAGAG |
All Microarray Probes Designed to Gene DMG400031065
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40901_PI426222305 | JHI_St_60k_v1 | DMT400079774 | AAGAGATGAATGCCATAACTACCCAATGTGAAACCGGAAATCCCATAGACGATTGCTGGC |
| CUST_40937_PI426222305 | JHI_St_60k_v1 | DMT400079776 | TAATCTCACACTTCCTCATCAACATCCTTACCCTGATGCTGTGGTTCAAGAAGTACAGAG |
Microarray Signals from CUST_40937_PI426222305
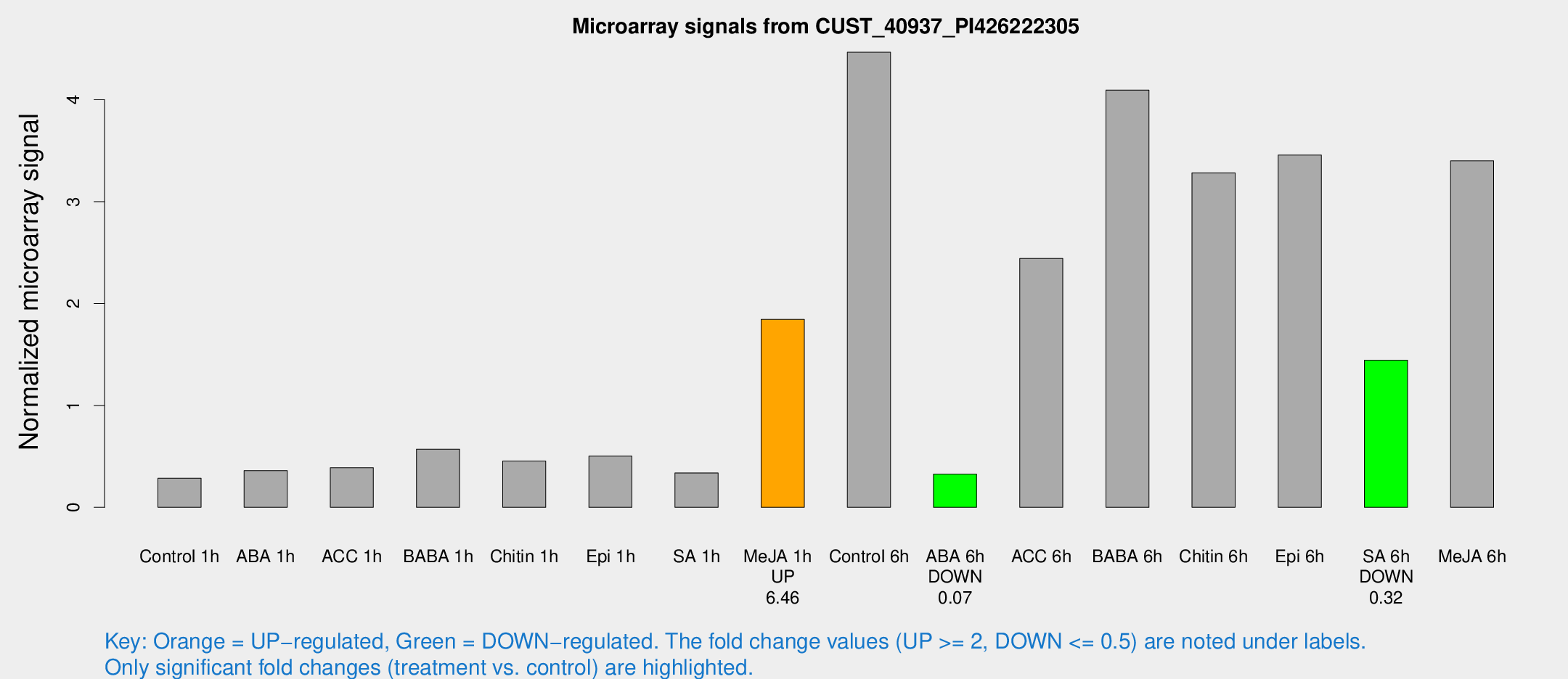
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.91025 | 3.43898 | 0.285445 | 0.165595 |
| ABA 1h | 6.78786 | 3.30089 | 0.360485 | 0.184778 |
| ACC 1h | 8.35495 | 4.37408 | 0.387908 | 0.200608 |
| BABA 1h | 12.4159 | 3.75471 | 0.571061 | 0.211354 |
| Chitin 1h | 9.77883 | 3.90896 | 0.453607 | 0.211588 |
| Epi 1h | 9.57975 | 3.36567 | 0.502072 | 0.209268 |
| SA 1h | 7.29811 | 3.45283 | 0.337217 | 0.167006 |
| Me-JA 1h | 31.1094 | 3.95357 | 1.8445 | 0.235284 |
| Control 6h | 97.5637 | 20.8635 | 4.46678 | 0.66117 |
| ABA 6h | 7.2533 | 3.6793 | 0.326015 | 0.170084 |
| ACC 6h | 67.8613 | 28.4777 | 2.44232 | 0.637073 |
| BABA 6h | 98.2265 | 19.349 | 4.09517 | 0.641785 |
| Chitin 6h | 78.4449 | 20.2441 | 3.28318 | 1.05231 |
| Epi 6h | 90.153 | 26.8909 | 3.45829 | 1.61373 |
| SA 6h | 34.2549 | 13.5702 | 1.44407 | 0.475058 |
| Me-JA 6h | 81.8442 | 28.7185 | 3.40099 | 1.36185 |
Source Transcript PGSC0003DMT400079776 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G04310.1 | +1 | 0.0 | 624 | 289/402 (72%) | Pectin lyase-like superfamily protein | chr5:1203356-1207352 REVERSE LENGTH=518 |
