Probe CUST_40282_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40282_PI426222305 | JHI_St_60k_v1 | DMT400036485 | GTTGGAAGGCAAAGCCATAGACCTTCTTCCTTAGATTTGGAAGATTATTTAAGGATTAAA |
All Microarray Probes Designed to Gene DMG400014054
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40276_PI426222305 | JHI_St_60k_v1 | DMT400036486 | GTTGGAAGGCAAAGCCATAGACCTTCTTCCTTAGATTTGGAAGATTATTTAAGGATTAAA |
| CUST_40282_PI426222305 | JHI_St_60k_v1 | DMT400036485 | GTTGGAAGGCAAAGCCATAGACCTTCTTCCTTAGATTTGGAAGATTATTTAAGGATTAAA |
Microarray Signals from CUST_40282_PI426222305
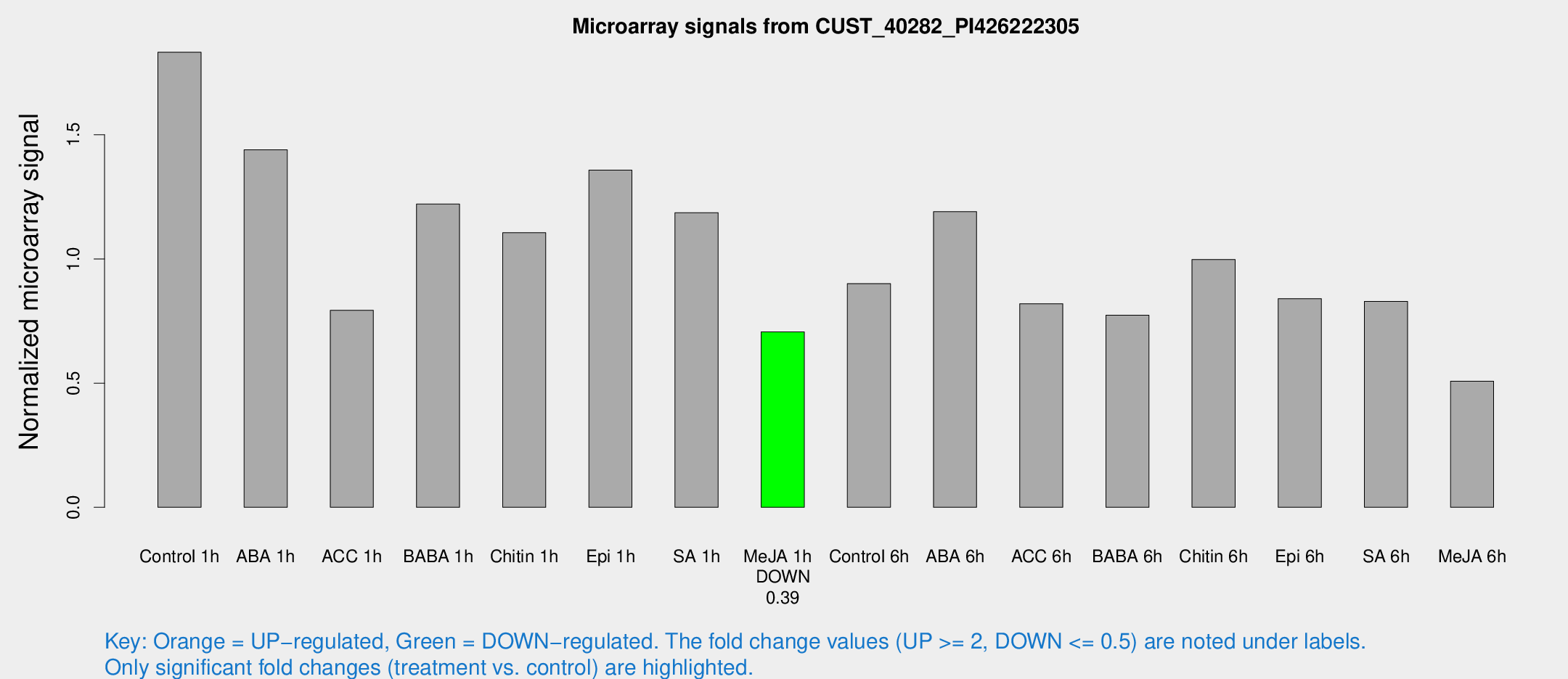
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2130.13 | 190.57 | 1.83238 | 0.105827 |
| ABA 1h | 1486.53 | 163.062 | 1.43916 | 0.0831432 |
| ACC 1h | 1021.66 | 259.622 | 0.79336 | 0.186651 |
| BABA 1h | 1427.38 | 305.173 | 1.22114 | 0.166185 |
| Chitin 1h | 1152.58 | 82.0888 | 1.10568 | 0.0639061 |
| Epi 1h | 1367.84 | 128.269 | 1.35738 | 0.123335 |
| SA 1h | 1410.78 | 81.773 | 1.18649 | 0.0685521 |
| Me-JA 1h | 665.841 | 38.6186 | 0.706182 | 0.0409098 |
| Control 6h | 1109.3 | 279.869 | 0.900496 | 0.178214 |
| ABA 6h | 1467.67 | 109.2 | 1.19031 | 0.068776 |
| ACC 6h | 1102.25 | 141.68 | 0.819735 | 0.0515312 |
| BABA 6h | 1066.28 | 287.347 | 0.773214 | 0.195473 |
| Chitin 6h | 1226.01 | 70.9006 | 0.997963 | 0.0576889 |
| Epi 6h | 1097.99 | 75.3862 | 0.840192 | 0.0587978 |
| SA 6h | 954.675 | 91.5705 | 0.828905 | 0.0479516 |
| Me-JA 6h | 625.175 | 170.588 | 0.508065 | 0.0946384 |
Source Transcript PGSC0003DMT400036485 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G40460.1 | +3 | 0.0 | 886 | 429/586 (73%) | Major facilitator superfamily protein | chr2:16897123-16901171 FORWARD LENGTH=583 |
