Probe CUST_4017_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4017_PI426222305 | JHI_St_60k_v1 | DMT400080788 | AAGTTTTATTCAAGGGCATCAAGCTTTAGTAGGGAAACGCAAGTGCCATGTTCTATCTAA |
All Microarray Probes Designed to Gene DMG400031467
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4017_PI426222305 | JHI_St_60k_v1 | DMT400080788 | AAGTTTTATTCAAGGGCATCAAGCTTTAGTAGGGAAACGCAAGTGCCATGTTCTATCTAA |
Microarray Signals from CUST_4017_PI426222305
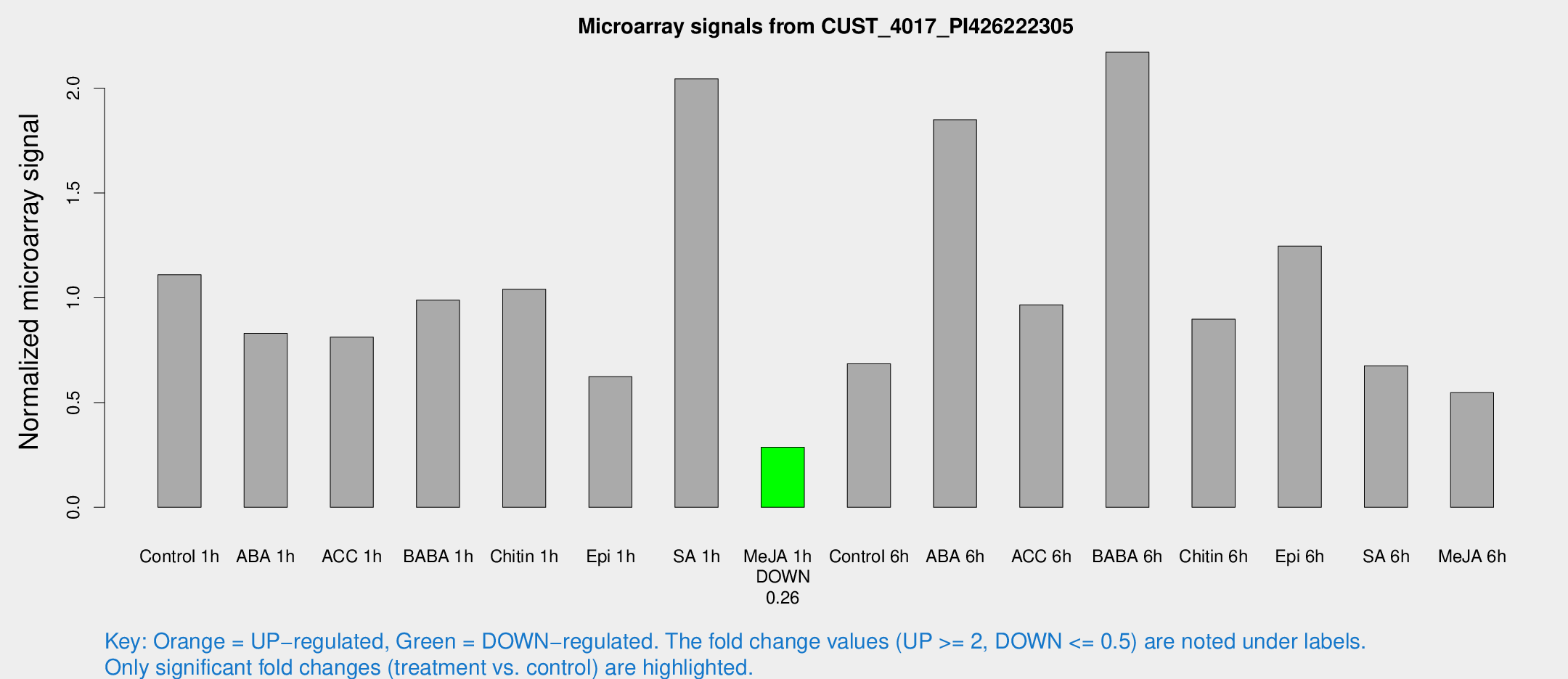
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 62.3082 | 17.4232 | 1.11014 | 0.253103 |
| ABA 1h | 44.5378 | 16.5251 | 0.829985 | 0.358297 |
| ACC 1h | 50.8207 | 19.2476 | 0.812472 | 0.304155 |
| BABA 1h | 49.6483 | 4.28759 | 0.988523 | 0.136943 |
| Chitin 1h | 48.784 | 4.12795 | 1.04092 | 0.087898 |
| Epi 1h | 28.6351 | 4.45133 | 0.623537 | 0.104025 |
| SA 1h | 109.596 | 9.68648 | 2.04488 | 0.131384 |
| Me-JA 1h | 12.2936 | 3.18139 | 0.28679 | 0.0762272 |
| Control 6h | 39.5615 | 11.314 | 0.685278 | 0.1866 |
| ABA 6h | 103.515 | 12.724 | 1.84941 | 0.169994 |
| ACC 6h | 58.5865 | 8.02246 | 0.965885 | 0.151657 |
| BABA 6h | 127.853 | 15.9166 | 2.17177 | 0.264162 |
| Chitin 6h | 50.8738 | 8.8027 | 0.897492 | 0.148225 |
| Epi 6h | 77.0543 | 19.3244 | 1.24688 | 0.305519 |
| SA 6h | 35.2385 | 4.78598 | 0.675034 | 0.174261 |
| Me-JA 6h | 29.5671 | 6.5976 | 0.546995 | 0.0754953 |
Source Transcript PGSC0003DMT400080788 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G13600.1 | +1 | 0.0 | 598 | 338/594 (57%) | calmodulin-binding family protein | chr3:4445102-4447383 FORWARD LENGTH=605 |
