Probe CUST_40092_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40092_PI426222305 | JHI_St_60k_v1 | DMT400015148 | CTACTCTACCATTTGAAGTCTTGTAGCAATGTTGTTATTGTTGCCAAAGTTGTGTAAATC |
All Microarray Probes Designed to Gene DMG400005915
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_40075_PI426222305 | JHI_St_60k_v1 | DMT400015147 | AATCTGCCTAGCATTGGATACGATACCACACCAAAGCAAGATGCTCCTATTTGGTACTAG |
| CUST_40092_PI426222305 | JHI_St_60k_v1 | DMT400015148 | CTACTCTACCATTTGAAGTCTTGTAGCAATGTTGTTATTGTTGCCAAAGTTGTGTAAATC |
Microarray Signals from CUST_40092_PI426222305
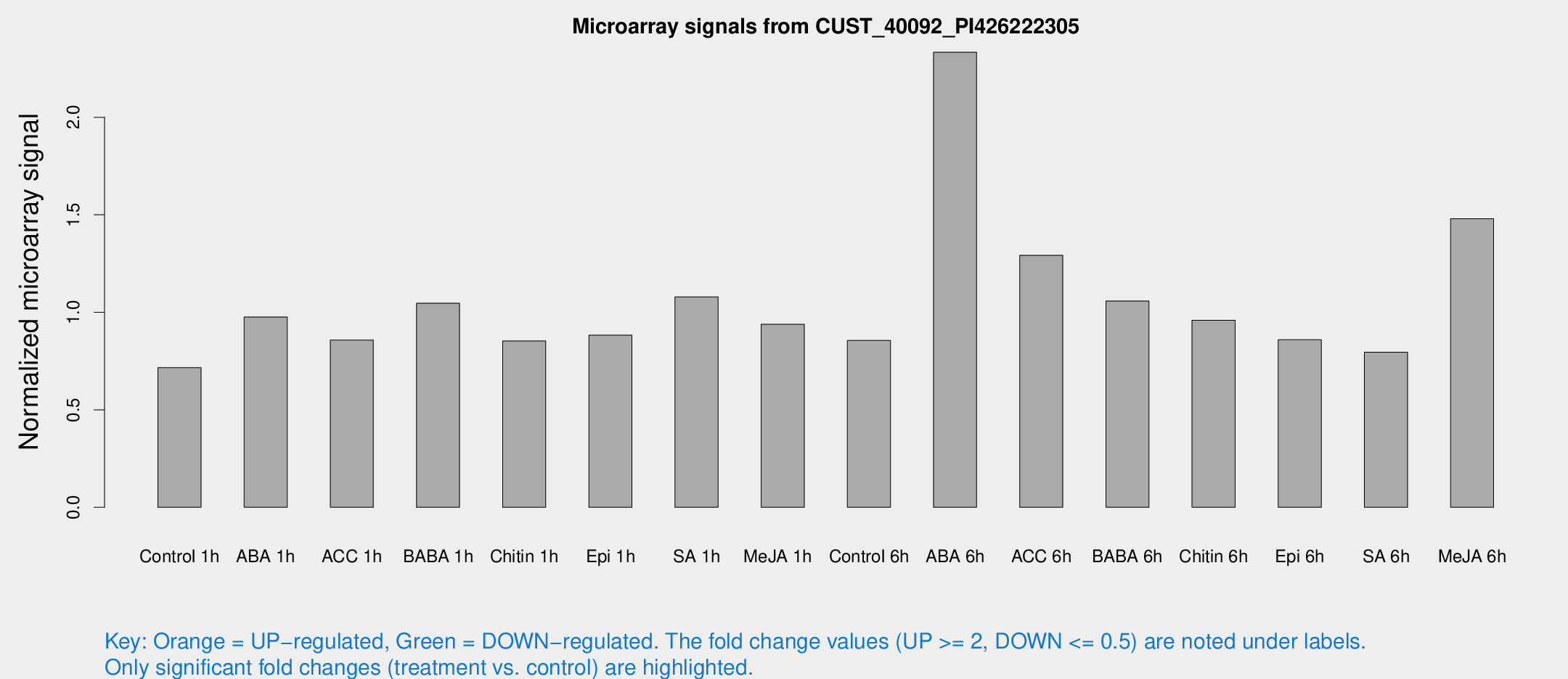
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1121.19 | 106.963 | 0.716178 | 0.081947 |
| ABA 1h | 1416.82 | 303.126 | 0.976099 | 0.198551 |
| ACC 1h | 1587.08 | 525.488 | 0.857395 | 0.323057 |
| BABA 1h | 1641.37 | 328.929 | 1.04652 | 0.130922 |
| Chitin 1h | 1220.64 | 206.87 | 0.85268 | 0.0881642 |
| Epi 1h | 1215.73 | 183.476 | 0.882789 | 0.136956 |
| SA 1h | 1779.85 | 316.429 | 1.07911 | 0.143465 |
| Me-JA 1h | 1210.34 | 165.85 | 0.938199 | 0.0542227 |
| Control 6h | 1399.6 | 288.168 | 0.855401 | 0.122888 |
| ABA 6h | 3969.91 | 719.692 | 2.33348 | 0.285522 |
| ACC 6h | 2313.83 | 194.116 | 1.2923 | 0.0747432 |
| BABA 6h | 1921.32 | 423.292 | 1.05784 | 0.188643 |
| Chitin 6h | 1583.25 | 91.4856 | 0.959231 | 0.05542 |
| Epi 6h | 1531.3 | 208.344 | 0.859091 | 0.0884825 |
| SA 6h | 1300.9 | 310.948 | 0.795458 | 0.126315 |
| Me-JA 6h | 2390.67 | 499.751 | 1.47965 | 0.195147 |
Source Transcript PGSC0003DMT400015148 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20770.1 | +2 | 0.0 | 657 | 359/624 (58%) | Ethylene insensitive 3 family protein | chr3:7260702-7262588 REVERSE LENGTH=628 |
