Probe CUST_39979_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39979_PI426222305 | JHI_St_60k_v1 | DMT400011155 | CCCATTTGATGAAGACTTGTAATTTGTTGACTCAATATTTCAATGGAAAAGCAGACCTCA |
All Microarray Probes Designed to Gene DMG400004367
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39979_PI426222305 | JHI_St_60k_v1 | DMT400011155 | CCCATTTGATGAAGACTTGTAATTTGTTGACTCAATATTTCAATGGAAAAGCAGACCTCA |
| CUST_39993_PI426222305 | JHI_St_60k_v1 | DMT400011154 | CTCGTGAAGATCATTTTGATCTCAATTTCTAGATTTACCTATTGCGAGGAGATCTTCACT |
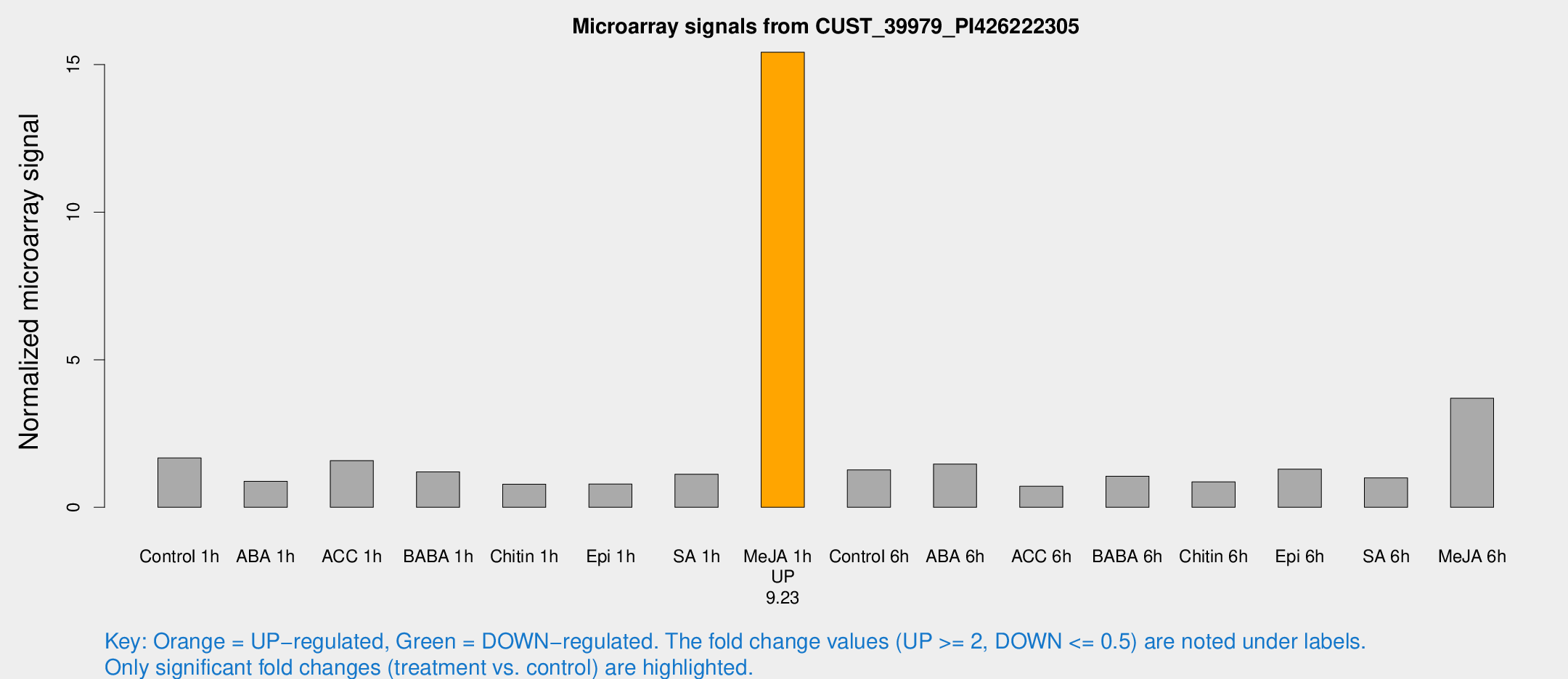
Microarray Signals from CUST_39979_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18.4764 | 6.23061 | 1.67036 | 0.796847 |
| ABA 1h | 7.72015 | 3.63148 | 0.878047 | 0.444847 |
| ACC 1h | 22.5262 | 13.89 | 1.58104 | 1.25151 |
| BABA 1h | 14.2557 | 7.5819 | 1.20048 | 0.641977 |
| Chitin 1h | 6.71401 | 3.72612 | 0.784364 | 0.436722 |
| Epi 1h | 6.47322 | 3.52876 | 0.789962 | 0.428255 |
| SA 1h | 11.4548 | 3.66223 | 1.12325 | 0.402596 |
| Me-JA 1h | 125.393 | 28.4204 | 15.4181 | 2.22157 |
| Control 6h | 15.0714 | 7.35136 | 1.27 | 0.620975 |
| ABA 6h | 16.738 | 5.12244 | 1.46697 | 0.723355 |
| ACC 6h | 7.94304 | 4.73548 | 0.713259 | 0.41341 |
| BABA 6h | 11.9359 | 4.30639 | 1.05486 | 0.428467 |
| Chitin 6h | 9.04864 | 4.20351 | 0.864289 | 0.433235 |
| Epi 6h | 15.0958 | 4.73864 | 1.29369 | 0.467833 |
| SA 6h | 9.48149 | 4.05136 | 0.995275 | 0.445167 |
| Me-JA 6h | 36.4583 | 7.19362 | 3.69378 | 1.17466 |
Source Transcript PGSC0003DMT400011155 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G19180.1 | +2 | 6e-23 | 95 | 71/219 (32%) | jasmonate-zim-domain protein 1 | chr1:6622312-6623271 FORWARD LENGTH=253 |
