Probe CUST_39941_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39941_PI426222305 | JHI_St_60k_v1 | DMT400076977 | CATTAGGTTCTGTGGCCAAATACCTTCAACTAGACAGAAAACAGGAAAAAGTGAATATTT |
All Microarray Probes Designed to Gene DMG400029941
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39935_PI426222305 | JHI_St_60k_v1 | DMT400076978 | TTGACAAGAATTTCTGAATGCTGAATAATTGGACAGTGCTGCTTTCCGCGAAACCTTGCT |
| CUST_39941_PI426222305 | JHI_St_60k_v1 | DMT400076977 | CATTAGGTTCTGTGGCCAAATACCTTCAACTAGACAGAAAACAGGAAAAAGTGAATATTT |
Microarray Signals from CUST_39941_PI426222305
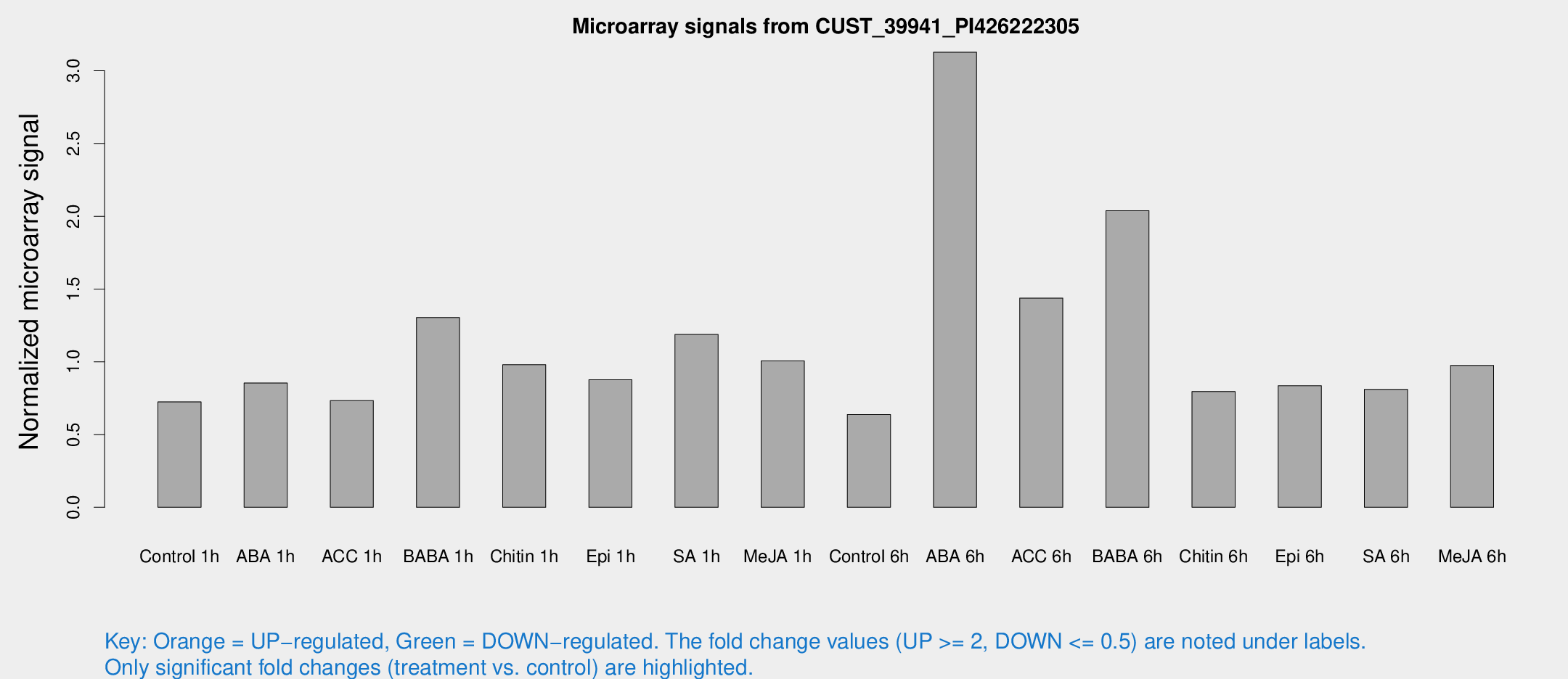
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 76.8884 | 9.39949 | 0.72479 | 0.072892 |
| ABA 1h | 79.5158 | 6.51356 | 0.854133 | 0.0635221 |
| ACC 1h | 111.703 | 46.1465 | 0.733371 | 0.616018 |
| BABA 1h | 138.979 | 34.0949 | 1.30387 | 0.230596 |
| Chitin 1h | 92.5438 | 6.49792 | 0.979575 | 0.0958227 |
| Epi 1h | 82.2082 | 14.269 | 0.8771 | 0.162233 |
| SA 1h | 132.704 | 26.7871 | 1.18766 | 0.164796 |
| Me-JA 1h | 88.3208 | 13.9433 | 1.00575 | 0.11481 |
| Control 6h | 87.6851 | 35.927 | 0.637841 | 0.383707 |
| ABA 6h | 351.048 | 33.9916 | 3.12725 | 0.209292 |
| ACC 6h | 183.948 | 48.5132 | 1.43776 | 0.195331 |
| BABA 6h | 253.242 | 65.0802 | 2.03863 | 0.444284 |
| Chitin 6h | 91.1898 | 15.3348 | 0.795977 | 0.128543 |
| Epi 6h | 99.56 | 10.3176 | 0.835262 | 0.0590559 |
| SA 6h | 88.7444 | 19.4107 | 0.810809 | 0.108494 |
| Me-JA 6h | 112.377 | 34.1015 | 0.974823 | 0.288283 |
Source Transcript PGSC0003DMT400076977 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G26070.1 | +3 | 3e-101 | 303 | 146/249 (59%) | Protein of unknown function (DUF778) | chr2:11105741-11106607 REVERSE LENGTH=250 |
