Probe CUST_39522_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39522_PI426222305 | JHI_St_60k_v1 | DMT400048729 | ATATGAGGCCAAAGTTGCTGCCAGGAAAATATTCAGAAATGTTGCCAAGCCTAGGTCCAA |
All Microarray Probes Designed to Gene DMG400018938
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39535_PI426222305 | JHI_St_60k_v1 | DMT400048728 | ATATGAGGCCAAAGTTGCTGCCAGGAAAATATTCAGAAATGTTGCCAAGCCTAGGTCCAA |
| CUST_39522_PI426222305 | JHI_St_60k_v1 | DMT400048729 | ATATGAGGCCAAAGTTGCTGCCAGGAAAATATTCAGAAATGTTGCCAAGCCTAGGTCCAA |
Microarray Signals from CUST_39522_PI426222305
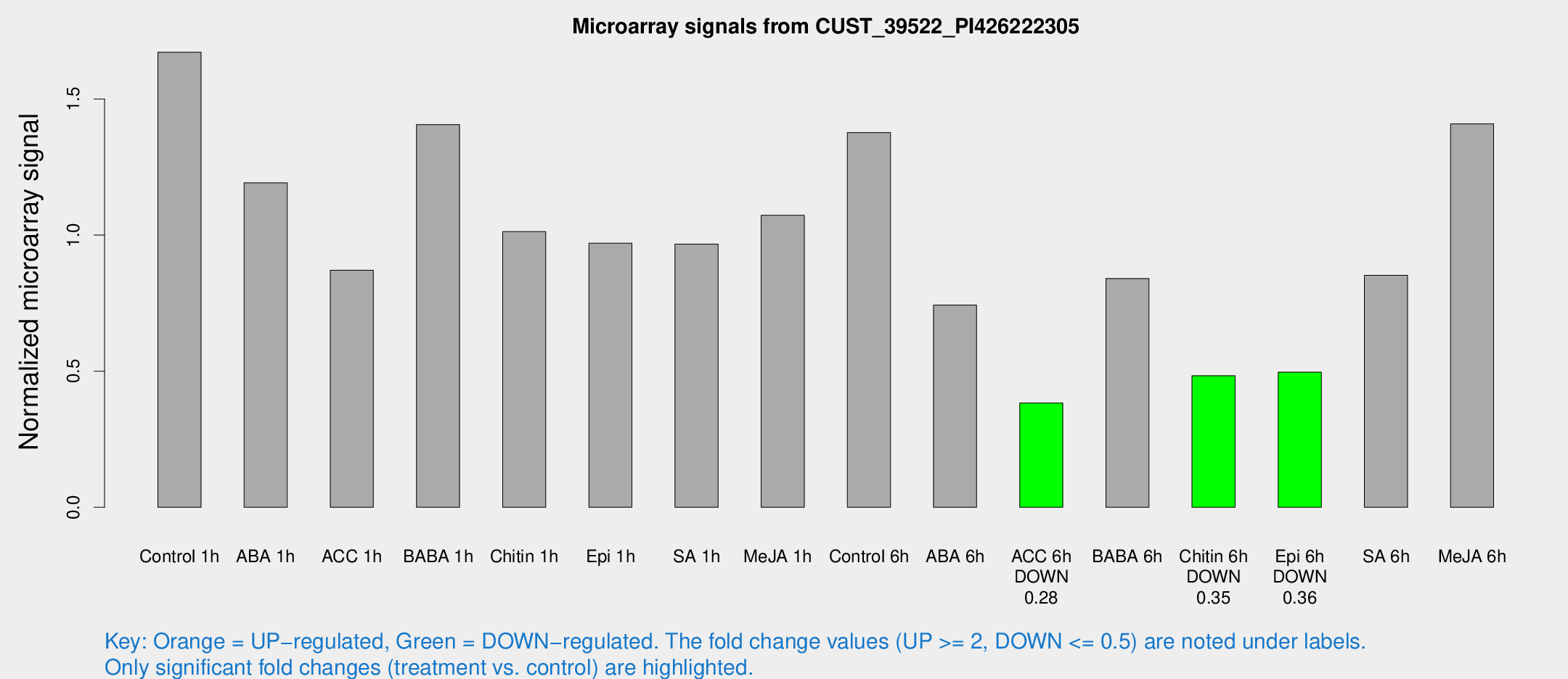
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 28.2683 | 5.24875 | 1.67194 | 0.27059 |
| ABA 1h | 17.688 | 3.54353 | 1.19196 | 0.260979 |
| ACC 1h | 15.264 | 4.50208 | 0.871032 | 0.282861 |
| BABA 1h | 24.3739 | 6.86371 | 1.40609 | 0.347228 |
| Chitin 1h | 15.7667 | 4.13802 | 1.01293 | 0.258265 |
| Epi 1h | 15.5117 | 4.65517 | 0.970339 | 0.403874 |
| SA 1h | 16.2838 | 3.64115 | 0.967169 | 0.221844 |
| Me-JA 1h | 16.8914 | 5.83086 | 1.07266 | 0.559012 |
| Control 6h | 23.34 | 4.97206 | 1.37692 | 0.255344 |
| ABA 6h | 14.9581 | 5.48655 | 0.742962 | 0.316964 |
| ACC 6h | 7.30127 | 4.33419 | 0.383222 | 0.222012 |
| BABA 6h | 17.7771 | 5.66899 | 0.840617 | 0.368808 |
| Chitin 6h | 8.49928 | 4.02084 | 0.483366 | 0.237392 |
| Epi 6h | 9.12178 | 4.37979 | 0.496338 | 0.234506 |
| SA 6h | 13.9297 | 3.96079 | 0.852183 | 0.254834 |
| Me-JA 6h | 23.092 | 3.86274 | 1.40885 | 0.242301 |
Source Transcript PGSC0003DMT400048729 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G78610.1 | +3 | 3e-132 | 424 | 246/463 (53%) | mechanosensitive channel of small conductance-like 6 | chr1:29569226-29572126 REVERSE LENGTH=856 |
