Probe CUST_39473_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39473_PI426222305 | JHI_St_60k_v1 | DMT400082120 | ATGTACCTTCATAAAGGTAGGAATCAAGCTGCGTACATACCATCCTCTCCAGTCAAGGTG |
All Microarray Probes Designed to Gene DMG400032271
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39473_PI426222305 | JHI_St_60k_v1 | DMT400082120 | ATGTACCTTCATAAAGGTAGGAATCAAGCTGCGTACATACCATCCTCTCCAGTCAAGGTG |
Microarray Signals from CUST_39473_PI426222305
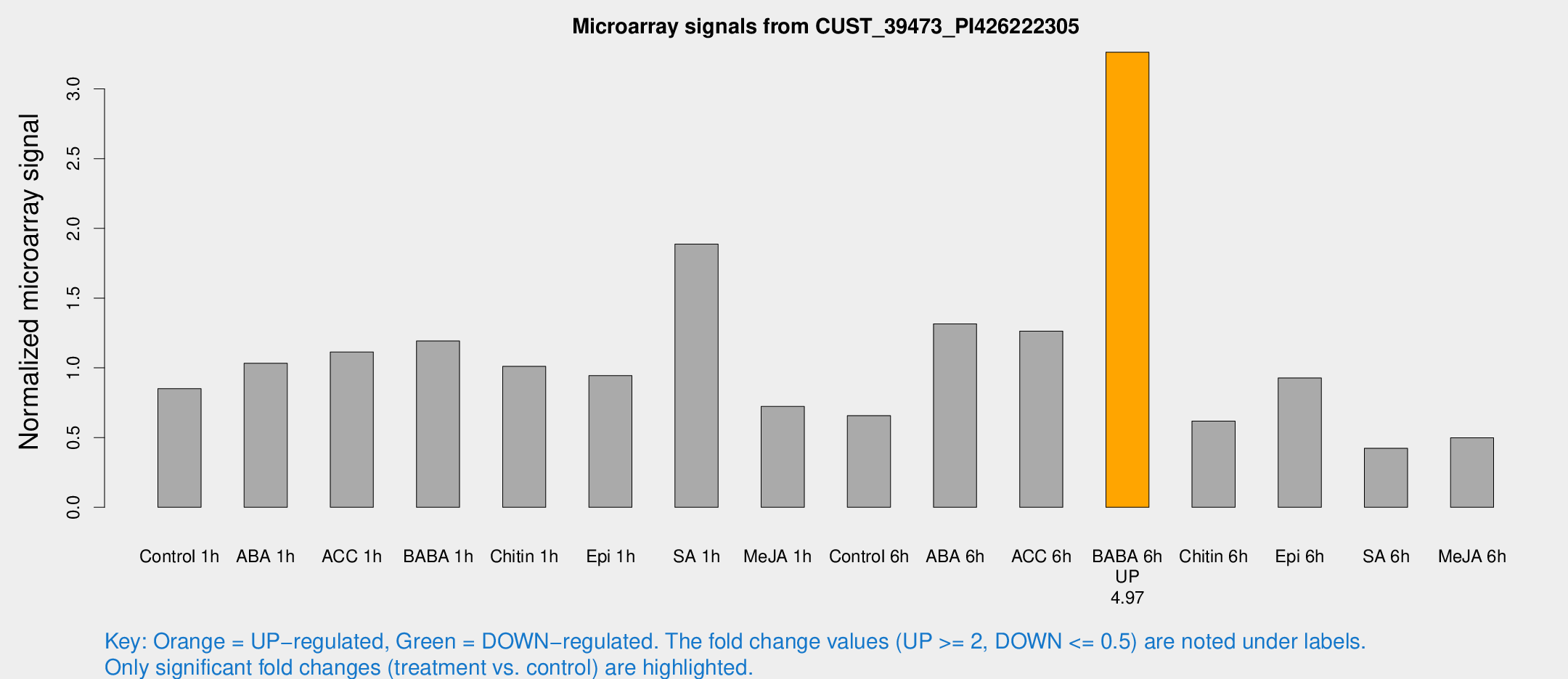
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 155.017 | 35.0231 | 0.850458 | 0.134795 |
| ABA 1h | 160.459 | 15.346 | 1.03346 | 0.164852 |
| ACC 1h | 215.733 | 53.9306 | 1.11361 | 0.259249 |
| BABA 1h | 204.444 | 29.2269 | 1.19273 | 0.0719401 |
| Chitin 1h | 160.057 | 17.5904 | 1.0103 | 0.0617648 |
| Epi 1h | 143.381 | 13.0478 | 0.944247 | 0.0949577 |
| SA 1h | 356.835 | 77.286 | 1.88683 | 0.367447 |
| Me-JA 1h | 104.972 | 16.051 | 0.723051 | 0.0539415 |
| Control 6h | 123.692 | 35.8694 | 0.657214 | 0.157125 |
| ABA 6h | 257.249 | 63.3166 | 1.31556 | 0.239119 |
| ACC 6h | 254.152 | 21.6193 | 1.26361 | 0.19936 |
| BABA 6h | 646.107 | 79.7059 | 3.26326 | 0.444258 |
| Chitin 6h | 115.818 | 11.5985 | 0.618564 | 0.0521935 |
| Epi 6h | 186.397 | 27.5639 | 0.927398 | 0.100609 |
| SA 6h | 73.3913 | 6.06857 | 0.42353 | 0.0591995 |
| Me-JA 6h | 108.177 | 52.2852 | 0.498612 | 0.215845 |
Source Transcript PGSC0003DMT400082120 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G32030.1 | +3 | 8e-61 | 192 | 96/161 (60%) | Acyl-CoA N-acyltransferases (NAT) superfamily protein | chr2:13632675-13633241 REVERSE LENGTH=188 |
