Probe CUST_39417_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39417_PI426222305 | JHI_St_60k_v1 | DMT400082122 | ATGCAGTGGTTACGGACCGACCATAGAAACCTATTAGCTATTTTATGTAAGTAAGTTTGT |
All Microarray Probes Designed to Gene DMG400032273
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39417_PI426222305 | JHI_St_60k_v1 | DMT400082122 | ATGCAGTGGTTACGGACCGACCATAGAAACCTATTAGCTATTTTATGTAAGTAAGTTTGT |
Microarray Signals from CUST_39417_PI426222305
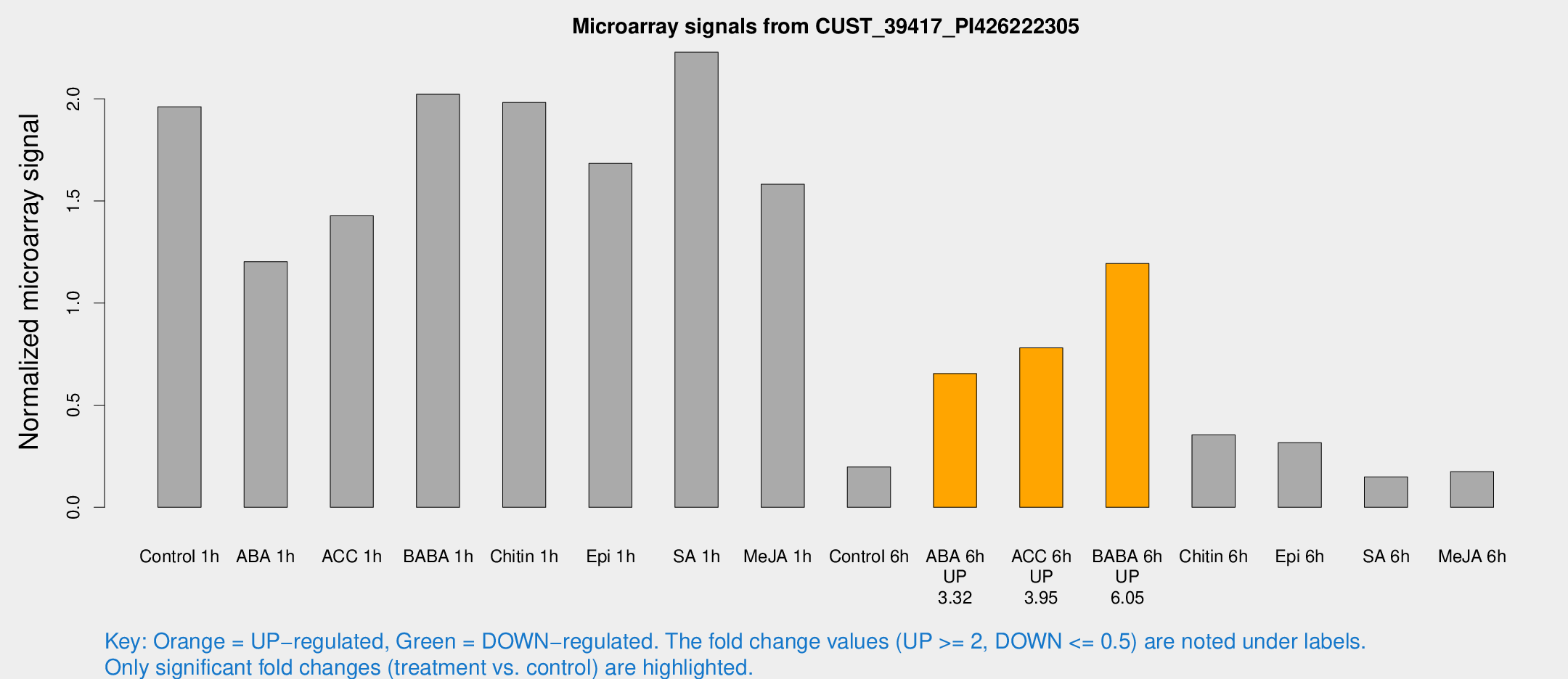
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3648.15 | 284.86 | 1.96156 | 0.113267 |
| ABA 1h | 2011.9 | 289 | 1.20265 | 0.180397 |
| ACC 1h | 3011.9 | 832.216 | 1.42748 | 0.406927 |
| BABA 1h | 3853.09 | 919.901 | 2.02259 | 0.346199 |
| Chitin 1h | 3299.36 | 190.74 | 1.98214 | 0.140935 |
| Epi 1h | 2697.44 | 155.939 | 1.6838 | 0.0972393 |
| SA 1h | 4270.3 | 386.567 | 2.22848 | 0.128675 |
| Me-JA 1h | 2447.48 | 399.533 | 1.58156 | 0.126243 |
| Control 6h | 388.699 | 95.6381 | 0.197448 | 0.0329684 |
| ABA 6h | 1376.42 | 316.061 | 0.655041 | 0.153396 |
| ACC 6h | 1683.89 | 208.188 | 0.780863 | 0.0972215 |
| BABA 6h | 2645.01 | 725.209 | 1.194 | 0.277961 |
| Chitin 6h | 703.425 | 61.1732 | 0.354577 | 0.0367547 |
| Epi 6h | 665.74 | 63.2014 | 0.316549 | 0.0419015 |
| SA 6h | 276.929 | 38.1194 | 0.148539 | 0.00976136 |
| Me-JA 6h | 364.753 | 130.251 | 0.174648 | 0.0499778 |
Source Transcript PGSC0003DMT400082122 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G32030.1 | +3 | 7e-64 | 202 | 97/164 (59%) | Acyl-CoA N-acyltransferases (NAT) superfamily protein | chr2:13632675-13633241 REVERSE LENGTH=188 |
