Probe CUST_39410_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39410_PI426222305 | JHI_St_60k_v1 | DMT400082121 | ATACCATAAGATGGACTGAACTCTCTGCTTTTTGAGACTAAAGCCAGAAGAAAGAAAAAA |
All Microarray Probes Designed to Gene DMG400032272
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39410_PI426222305 | JHI_St_60k_v1 | DMT400082121 | ATACCATAAGATGGACTGAACTCTCTGCTTTTTGAGACTAAAGCCAGAAGAAAGAAAAAA |
Microarray Signals from CUST_39410_PI426222305
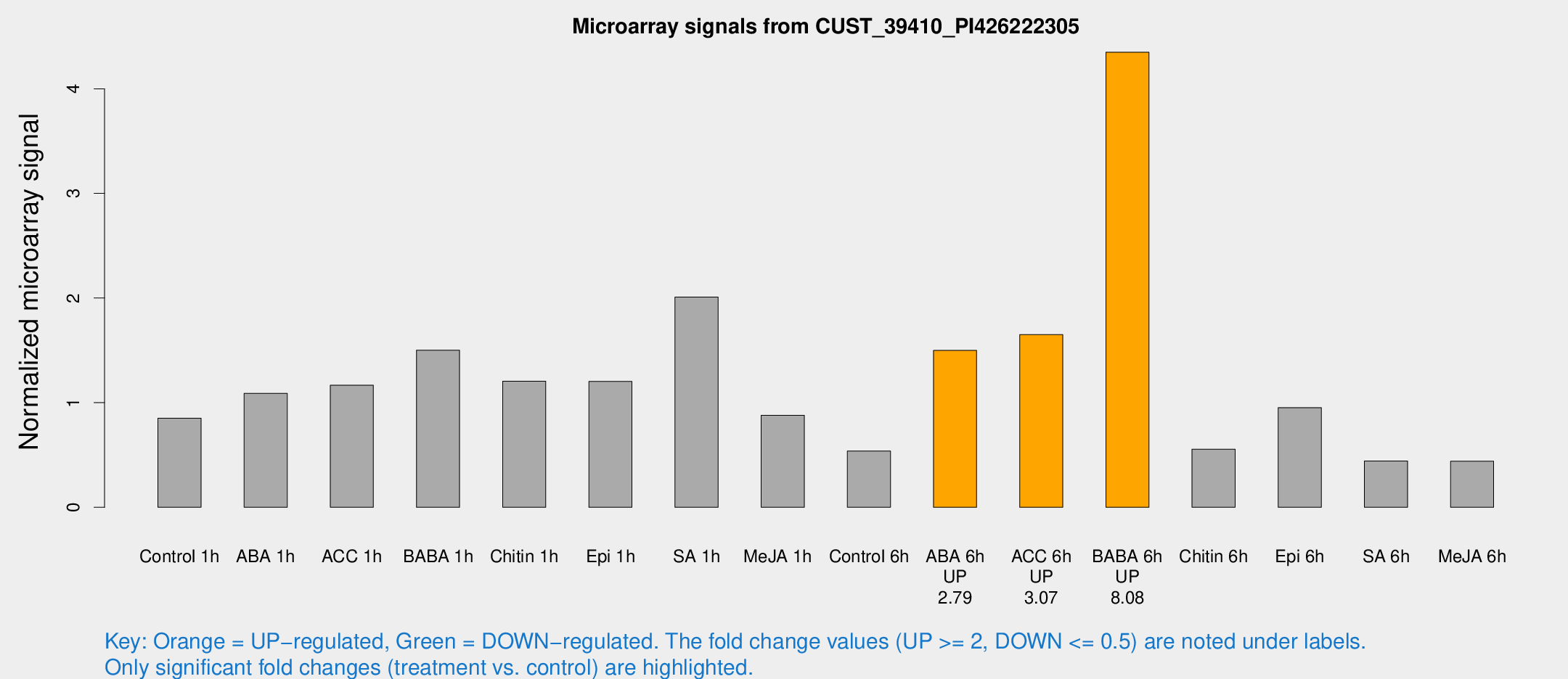
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 305.381 | 46.6805 | 0.851334 | 0.0721021 |
| ABA 1h | 344.521 | 48.4712 | 1.08958 | 0.217857 |
| ACC 1h | 457.119 | 117.45 | 1.16684 | 0.279308 |
| BABA 1h | 523.226 | 91.2494 | 1.50108 | 0.155798 |
| Chitin 1h | 395.327 | 80.8285 | 1.20584 | 0.15917 |
| Epi 1h | 368.195 | 33.0068 | 1.20273 | 0.130925 |
| SA 1h | 791.174 | 205.838 | 2.00948 | 0.567227 |
| Me-JA 1h | 252.84 | 15.9304 | 0.878913 | 0.0520257 |
| Control 6h | 201.234 | 52.2798 | 0.538055 | 0.102371 |
| ABA 6h | 597.808 | 152.622 | 1.49969 | 0.313405 |
| ACC 6h | 679.004 | 97.5757 | 1.65075 | 0.352036 |
| BABA 6h | 1753.98 | 266.401 | 4.34937 | 0.785762 |
| Chitin 6h | 210.703 | 26.0411 | 0.555832 | 0.0560658 |
| Epi 6h | 384.075 | 51.0681 | 0.952962 | 0.0969716 |
| SA 6h | 153.958 | 10.1549 | 0.44169 | 0.0482538 |
| Me-JA 6h | 196.097 | 100.104 | 0.440614 | 0.20551 |
Source Transcript PGSC0003DMT400082121 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G32030.1 | +3 | 2e-59 | 192 | 97/178 (54%) | Acyl-CoA N-acyltransferases (NAT) superfamily protein | chr2:13632675-13633241 REVERSE LENGTH=188 |
