Probe CUST_39320_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39320_PI426222305 | JHI_St_60k_v1 | DMT400012770 | ATTAAAGTCCGGTTTCTTCTTGTTGAATACAGTTTCTTGATGAGAAGGATCGTGTGGGTC |
All Microarray Probes Designed to Gene DMG400004977
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_39328_PI426222305 | JHI_St_60k_v1 | DMT400012771 | TTAGCTTAGTTTGTGAGATTATGCAACTTGAGTTTCTTGATGAGAAGGATCGTGTGGGTC |
| CUST_39320_PI426222305 | JHI_St_60k_v1 | DMT400012770 | ATTAAAGTCCGGTTTCTTCTTGTTGAATACAGTTTCTTGATGAGAAGGATCGTGTGGGTC |
Microarray Signals from CUST_39320_PI426222305
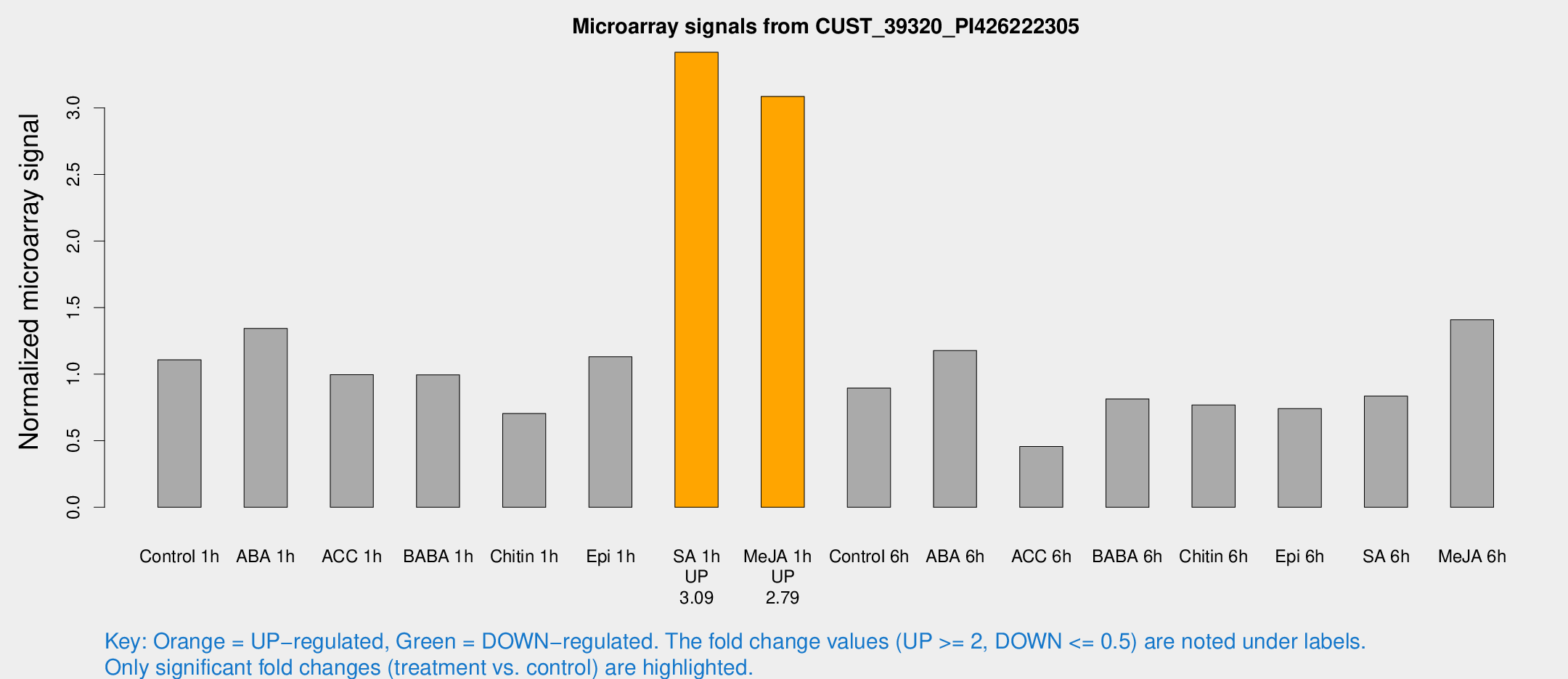
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 135.011 | 22.8863 | 1.10738 | 0.110439 |
| ABA 1h | 144.115 | 21.5408 | 1.34265 | 0.18165 |
| ACC 1h | 123.487 | 15.3342 | 0.996619 | 0.0647216 |
| BABA 1h | 116.236 | 16.1081 | 0.995181 | 0.0646173 |
| Chitin 1h | 75.5148 | 5.37626 | 0.704399 | 0.05063 |
| Epi 1h | 118.376 | 16.348 | 1.13048 | 0.157586 |
| SA 1h | 421.487 | 47.0965 | 3.41758 | 0.360373 |
| Me-JA 1h | 301.062 | 26.7922 | 3.08551 | 0.180978 |
| Control 6h | 114.817 | 29.4122 | 0.895461 | 0.175376 |
| ABA 6h | 153.651 | 27.3573 | 1.17734 | 0.161369 |
| ACC 6h | 62.2149 | 5.23506 | 0.456256 | 0.0860559 |
| BABA 6h | 108.75 | 8.56901 | 0.814035 | 0.0538044 |
| Chitin 6h | 97.9357 | 9.60412 | 0.768194 | 0.0523076 |
| Epi 6h | 101.898 | 17.533 | 0.74139 | 0.13095 |
| SA 6h | 114.109 | 43.9137 | 0.834747 | 0.266962 |
| Me-JA 6h | 170.824 | 27.75 | 1.40834 | 0.112153 |
Source Transcript PGSC0003DMT400012770 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G41210.1 | +1 | 5e-98 | 295 | 147/284 (52%) | glutathione S-transferase THETA 1 | chr5:16492550-16493884 REVERSE LENGTH=245 |
