Probe CUST_38957_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38957_PI426222305 | JHI_St_60k_v1 | DMT400070252 | TCTTTCGCATTTGAGTCTAGGAGTTTCTCATGACTTCCATGAATTGAATCATTTTGCTAG |
All Microarray Probes Designed to Gene DMG400027316
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38955_PI426222305 | JHI_St_60k_v1 | DMT400070251 | TCTTTCGCATTTGAGTCTAGGAGTTTCTCATGACTTCCATGAATTGAATCATTTTGCTAG |
| CUST_38957_PI426222305 | JHI_St_60k_v1 | DMT400070252 | TCTTTCGCATTTGAGTCTAGGAGTTTCTCATGACTTCCATGAATTGAATCATTTTGCTAG |
Microarray Signals from CUST_38957_PI426222305
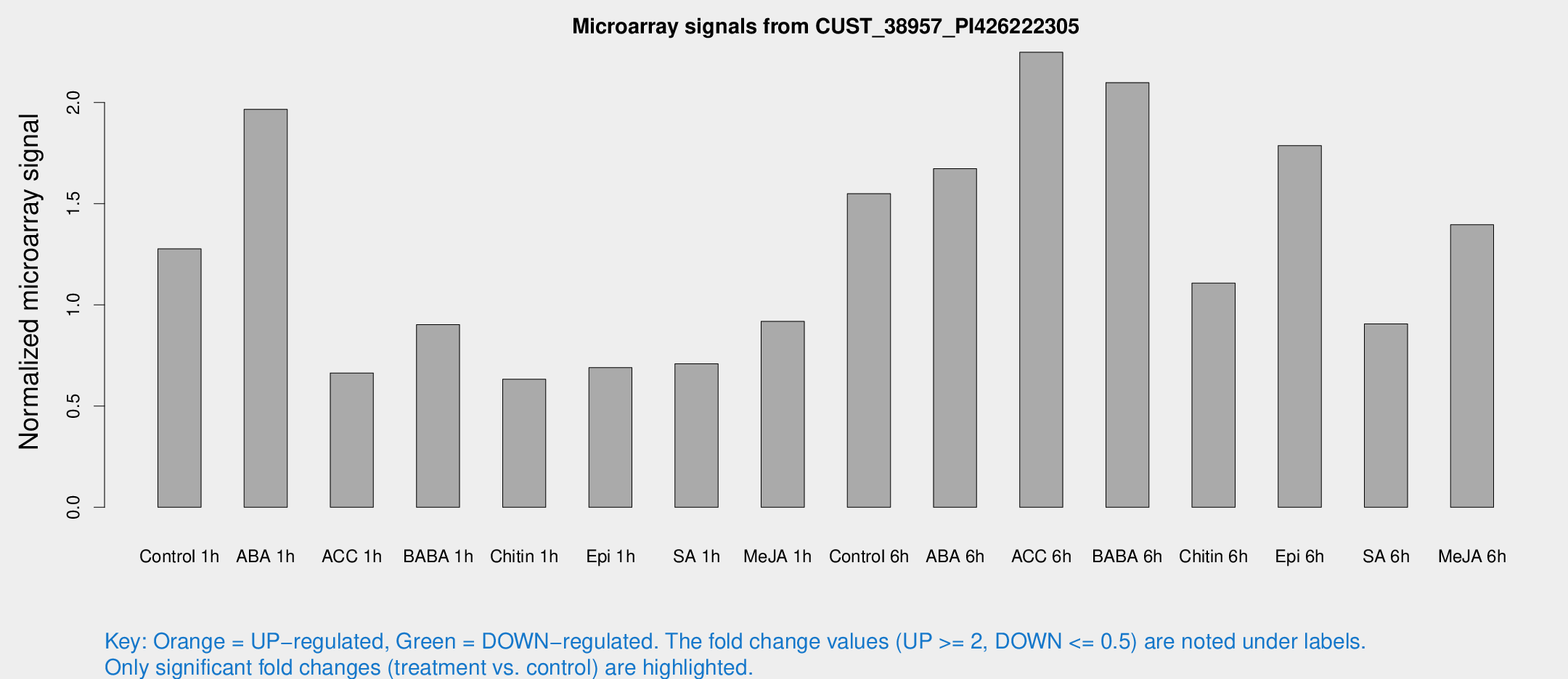
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.9683 | 6.02356 | 1.27692 | 0.39455 |
| ABA 1h | 19.9675 | 4.32669 | 1.96627 | 0.445859 |
| ACC 1h | 7.59154 | 4.46254 | 0.663756 | 0.385337 |
| BABA 1h | 10.0763 | 3.93657 | 0.902646 | 0.382877 |
| Chitin 1h | 6.29703 | 3.67465 | 0.632514 | 0.366629 |
| Epi 1h | 6.5736 | 3.63619 | 0.689919 | 0.377766 |
| SA 1h | 8.19067 | 3.70745 | 0.709209 | 0.33238 |
| Me-JA 1h | 8.57914 | 3.85439 | 0.918345 | 0.448668 |
| Control 6h | 21.345 | 9.36266 | 1.5495 | 0.738075 |
| ABA 6h | 24.4085 | 8.8161 | 1.67323 | 0.972102 |
| ACC 6h | 28.6518 | 4.76561 | 2.24847 | 0.376955 |
| BABA 6h | 29.2677 | 8.76104 | 2.09801 | 0.784476 |
| Chitin 6h | 16.8981 | 8.97118 | 1.10764 | 0.717992 |
| Epi 6h | 25.2446 | 9.59304 | 1.78698 | 0.683794 |
| SA 6h | 10.2701 | 4.05651 | 0.906234 | 0.39725 |
| Me-JA 6h | 21.1643 | 11.9429 | 1.39596 | 0.899724 |
Source Transcript PGSC0003DMT400070252 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G57810.2 | +2 | 2e-53 | 188 | 85/139 (61%) | Cysteine proteinases superfamily protein | chr3:21416336-21417809 FORWARD LENGTH=317 |
