Probe CUST_38270_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38270_PI426222305 | JHI_St_60k_v1 | DMT400067234 | TAATAATGCAGAGATTAGTAGTGCAGGGATAAATAATGCAGGGTCAGTTAGGGTATATTT |
All Microarray Probes Designed to Gene DMG400026148
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38270_PI426222305 | JHI_St_60k_v1 | DMT400067234 | TAATAATGCAGAGATTAGTAGTGCAGGGATAAATAATGCAGGGTCAGTTAGGGTATATTT |
Microarray Signals from CUST_38270_PI426222305
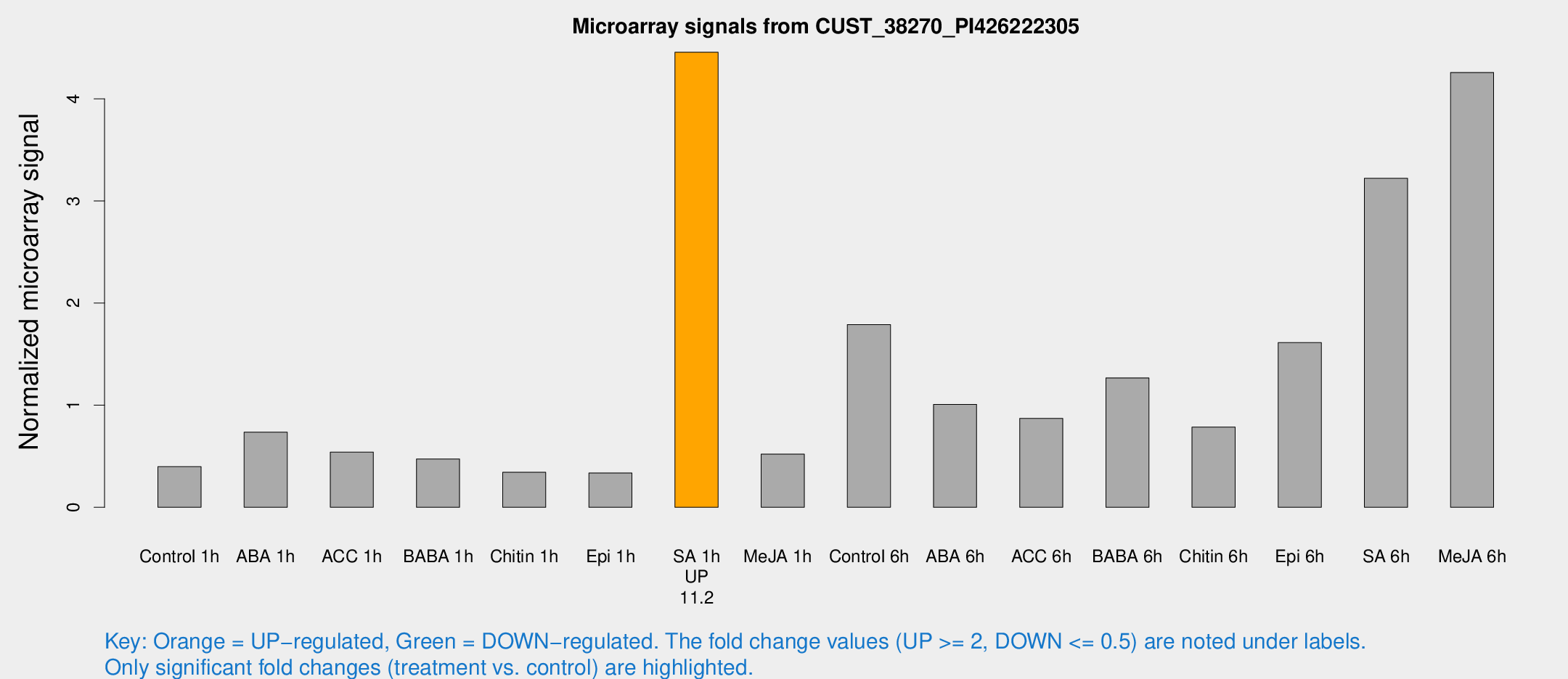
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 165.691 | 32.5335 | 0.397765 | 0.0990466 |
| ABA 1h | 266.407 | 46.4635 | 0.735797 | 0.125143 |
| ACC 1h | 252.941 | 98.4378 | 0.539956 | 0.200408 |
| BABA 1h | 206.237 | 68.1343 | 0.473482 | 0.138159 |
| Chitin 1h | 125.112 | 18.0856 | 0.342512 | 0.0229843 |
| Epi 1h | 120.967 | 27.1349 | 0.335599 | 0.0753693 |
| SA 1h | 1888.36 | 340.152 | 4.45596 | 0.71115 |
| Me-JA 1h | 182.973 | 45.3043 | 0.520874 | 0.126803 |
| Control 6h | 751.888 | 177.31 | 1.7895 | 0.28304 |
| ABA 6h | 428.665 | 32.7765 | 1.00762 | 0.0589707 |
| ACC 6h | 438.872 | 132.97 | 0.870426 | 0.467511 |
| BABA 6h | 572.28 | 68.5681 | 1.26713 | 0.194212 |
| Chitin 6h | 348.616 | 70.5513 | 0.786338 | 0.208582 |
| Epi 6h | 728.494 | 62.4802 | 1.61211 | 0.0936241 |
| SA 6h | 1413.25 | 395.633 | 3.22156 | 0.791086 |
| Me-JA 6h | 1899.35 | 550.52 | 4.25788 | 1.87219 |
Source Transcript PGSC0003DMT400067234 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G11360.1 | +2 | 2e-60 | 197 | 115/184 (63%) | Adenine nucleotide alpha hydrolases-like superfamily protein | chr1:3822171-3822899 REVERSE LENGTH=242 |
