Probe CUST_38260_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38260_PI426222305 | JHI_St_60k_v1 | DMT400067341 | GACATTTACTTTGCCAGTACCTAGATGAGAGGATTTAGCTTTCCATATTTCTTATGGAAT |
All Microarray Probes Designed to Gene DMG400026183
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38281_PI426222305 | JHI_St_60k_v1 | DMT400067343 | GACATTTACTTTGCCAGTACCTAGATGAGAGGATTTAGCTTTCCATATTTCTTATGGAAT |
| CUST_38260_PI426222305 | JHI_St_60k_v1 | DMT400067341 | GACATTTACTTTGCCAGTACCTAGATGAGAGGATTTAGCTTTCCATATTTCTTATGGAAT |
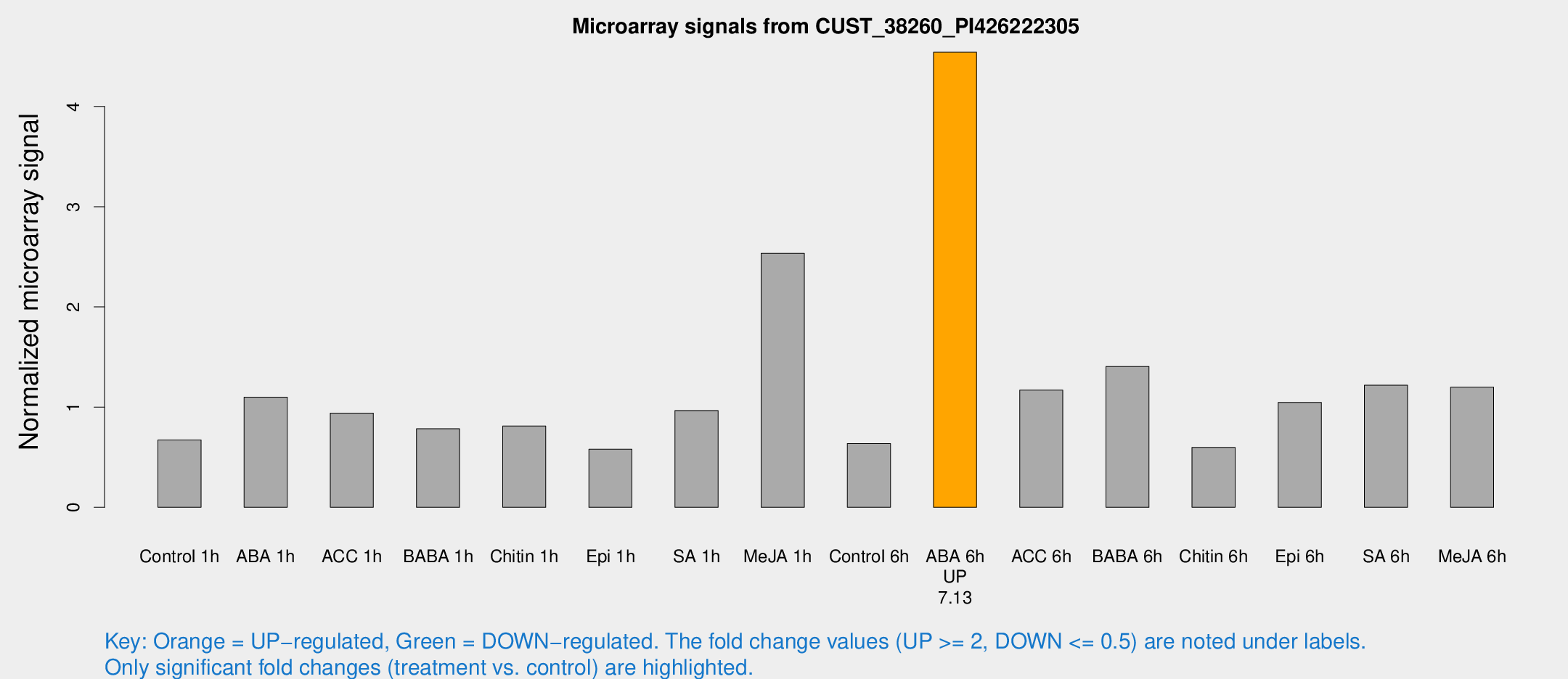
Microarray Signals from CUST_38260_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.2869 | 3.37274 | 0.673008 | 0.242076 |
| ABA 1h | 16.3498 | 4.59581 | 1.09926 | 0.354494 |
| ACC 1h | 17.9448 | 7.40561 | 0.93938 | 0.444599 |
| BABA 1h | 13.0168 | 4.07089 | 0.783565 | 0.274415 |
| Chitin 1h | 12.333 | 3.49997 | 0.811546 | 0.33724 |
| Epi 1h | 7.88625 | 3.09353 | 0.578934 | 0.252178 |
| SA 1h | 16.8952 | 5.68396 | 0.965645 | 0.338733 |
| Me-JA 1h | 33.7186 | 9.68619 | 2.53496 | 0.553117 |
| Control 6h | 10.2128 | 3.34391 | 0.636727 | 0.24408 |
| ABA 6h | 74.0649 | 9.55257 | 4.54205 | 0.613137 |
| ACC 6h | 21.979 | 5.5824 | 1.16911 | 0.58989 |
| BABA 6h | 25.5846 | 6.89458 | 1.40583 | 0.387072 |
| Chitin 6h | 10.3222 | 3.63105 | 0.597486 | 0.24734 |
| Epi 6h | 18.9059 | 4.77414 | 1.0463 | 0.286717 |
| SA 6h | 19.4969 | 5.01758 | 1.21812 | 0.510204 |
| Me-JA 6h | 20.0995 | 5.91397 | 1.19866 | 0.334275 |
Source Transcript PGSC0003DMT400067341 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G21390.1 | +1 | 2e-160 | 241 | 151/448 (34%) | S-locus lectin protein kinase family protein | chr4:11394458-11397474 REVERSE LENGTH=849 |
