Probe CUST_38230_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38230_PI426222305 | JHI_St_60k_v1 | DMT400067360 | ATTAACATGGAAAGGTCATAACTCAGTCTACGCAACATCTTGGGTCATAGTCCCCACCAC |
All Microarray Probes Designed to Gene DMG400026191
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38230_PI426222305 | JHI_St_60k_v1 | DMT400067360 | ATTAACATGGAAAGGTCATAACTCAGTCTACGCAACATCTTGGGTCATAGTCCCCACCAC |
Microarray Signals from CUST_38230_PI426222305
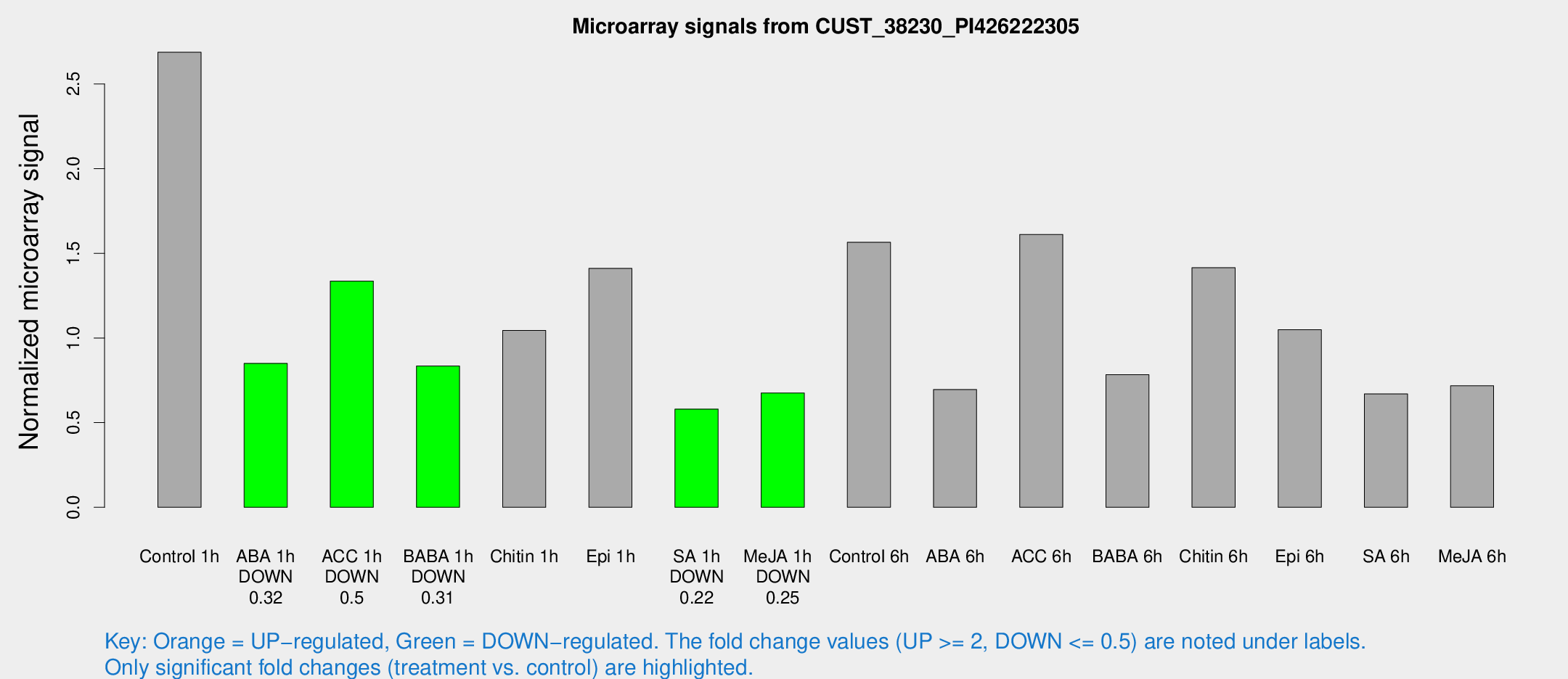
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 27.1019 | 3.46363 | 2.68757 | 0.356396 |
| ABA 1h | 7.87595 | 3.01487 | 0.849779 | 0.35988 |
| ACC 1h | 13.7075 | 3.73095 | 1.33599 | 0.358185 |
| BABA 1h | 8.5051 | 3.28126 | 0.834654 | 0.359768 |
| Chitin 1h | 10.4757 | 3.5144 | 1.04444 | 0.395396 |
| Epi 1h | 12.2507 | 3.11703 | 1.41095 | 0.363995 |
| SA 1h | 5.97461 | 3.11379 | 0.5802 | 0.307727 |
| Me-JA 1h | 5.49456 | 3.18191 | 0.674773 | 0.389535 |
| Control 6h | 16.4712 | 3.95966 | 1.5655 | 0.35546 |
| ABA 6h | 7.81682 | 3.29025 | 0.695774 | 0.335069 |
| ACC 6h | 24.1466 | 9.51528 | 1.61142 | 1.08059 |
| BABA 6h | 8.88479 | 3.56617 | 0.783199 | 0.332874 |
| Chitin 6h | 15.5133 | 3.61997 | 1.4151 | 0.365887 |
| Epi 6h | 13.2524 | 3.84235 | 1.04877 | 0.382325 |
| SA 6h | 6.64009 | 3.35302 | 0.669546 | 0.34217 |
| Me-JA 6h | 7.37979 | 3.19682 | 0.717941 | 0.333484 |
Source Transcript PGSC0003DMT400067360 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G49950.1 | +2 | 1e-114 | 348 | 189/404 (47%) | GRAS family transcription factor | chr3:18522570-18523802 FORWARD LENGTH=410 |
