Probe CUST_38228_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38228_PI426222305 | JHI_St_60k_v1 | DMT400067337 | GATAAGAGTTGTTGAATTTGTTCTAGTACTACTAATCTTCTTAGGGATAGGACCATTTGC |
All Microarray Probes Designed to Gene DMG400026181
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38228_PI426222305 | JHI_St_60k_v1 | DMT400067337 | GATAAGAGTTGTTGAATTTGTTCTAGTACTACTAATCTTCTTAGGGATAGGACCATTTGC |
| CUST_38273_PI426222305 | JHI_St_60k_v1 | DMT400067336 | GAAGCTGATCCGGATCTTCATTCAGATGTATGAACTTTTCCTACCTTTTCTTATTCTTGT |
Microarray Signals from CUST_38228_PI426222305
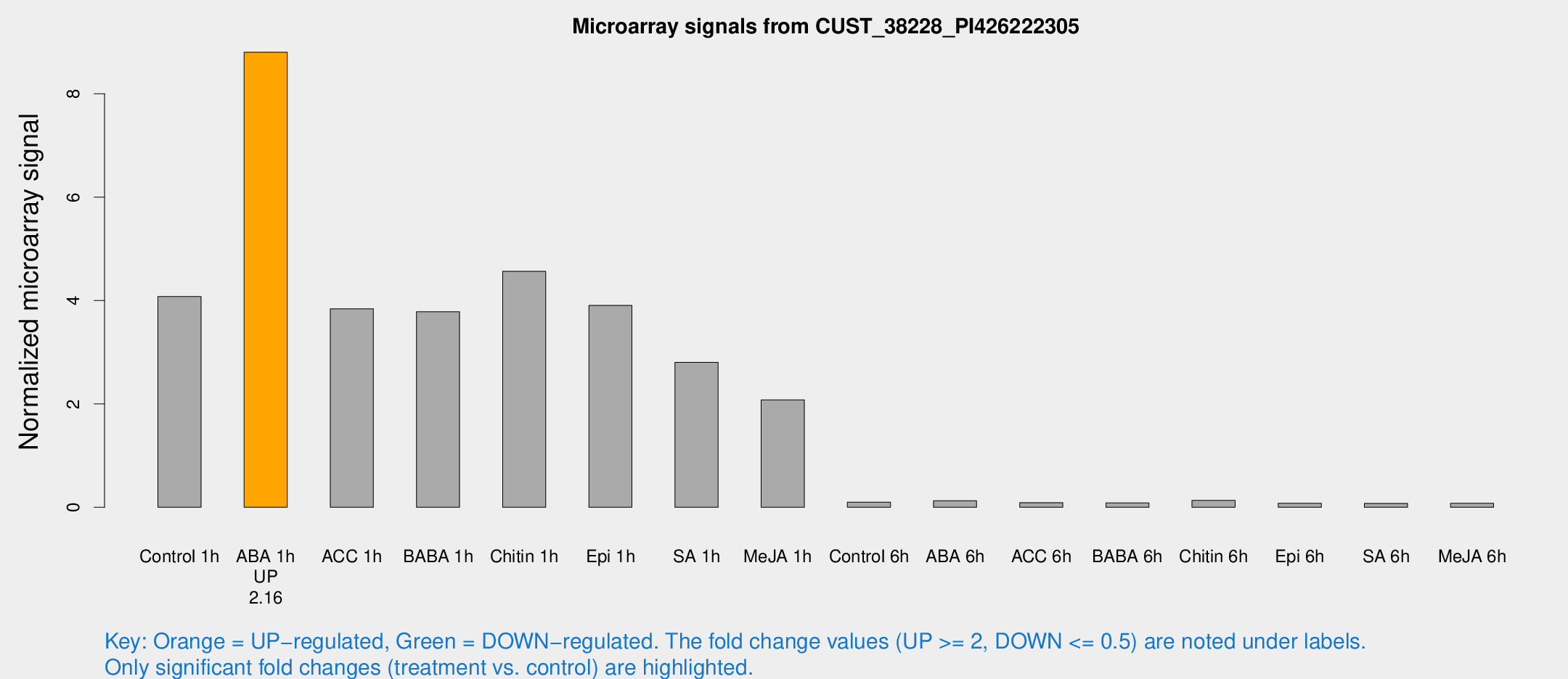
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9727.5 | 1067.61 | 4.07726 | 0.284746 |
| ABA 1h | 18665.1 | 2438.6 | 8.80466 | 1.00773 |
| ACC 1h | 9914.63 | 2255.3 | 3.8406 | 0.755847 |
| BABA 1h | 9132.4 | 2041 | 3.78363 | 0.596796 |
| Chitin 1h | 9759.44 | 945.675 | 4.5654 | 0.263588 |
| Epi 1h | 7971.16 | 460.935 | 3.90545 | 0.243466 |
| SA 1h | 7168.11 | 1658.41 | 2.80335 | 0.503396 |
| Me-JA 1h | 4101.73 | 658.825 | 2.07978 | 0.152389 |
| Control 6h | 245.402 | 48.6876 | 0.100244 | 0.0202731 |
| ABA 6h | 317.396 | 26.2524 | 0.12595 | 0.0144631 |
| ACC 6h | 247.324 | 35.861 | 0.0894255 | 0.0168212 |
| BABA 6h | 224.242 | 20.7773 | 0.0843858 | 0.00739124 |
| Chitin 6h | 334.077 | 19.6987 | 0.133237 | 0.00785423 |
| Epi 6h | 207.069 | 12.722 | 0.0777494 | 0.0104819 |
| SA 6h | 184.173 | 35.1453 | 0.0760451 | 0.00756378 |
| Me-JA 6h | 182.772 | 23.4022 | 0.0767441 | 0.00469937 |
Source Transcript PGSC0003DMT400067337 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27310.1 | +3 | 8e-31 | 118 | 50/72 (69%) | B-box type zinc finger family protein | chr4:13675853-13676616 FORWARD LENGTH=223 |
