Probe CUST_38119_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38119_PI426222305 | JHI_St_60k_v1 | DMT400054793 | ACATGATTCTTCTGAGGATCTGAGTAACATTGTTGATCTGTAGACGAAGACGGAGATGTA |
All Microarray Probes Designed to Gene DMG400021265
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_38119_PI426222305 | JHI_St_60k_v1 | DMT400054793 | ACATGATTCTTCTGAGGATCTGAGTAACATTGTTGATCTGTAGACGAAGACGGAGATGTA |
| CUST_38195_PI426222305 | JHI_St_60k_v1 | DMT400054792 | ACATGATTCTTCTGAGGATCTGAGTAACATTGTTGATCTGTAGACGAAGACGGAGATGTA |
Microarray Signals from CUST_38119_PI426222305
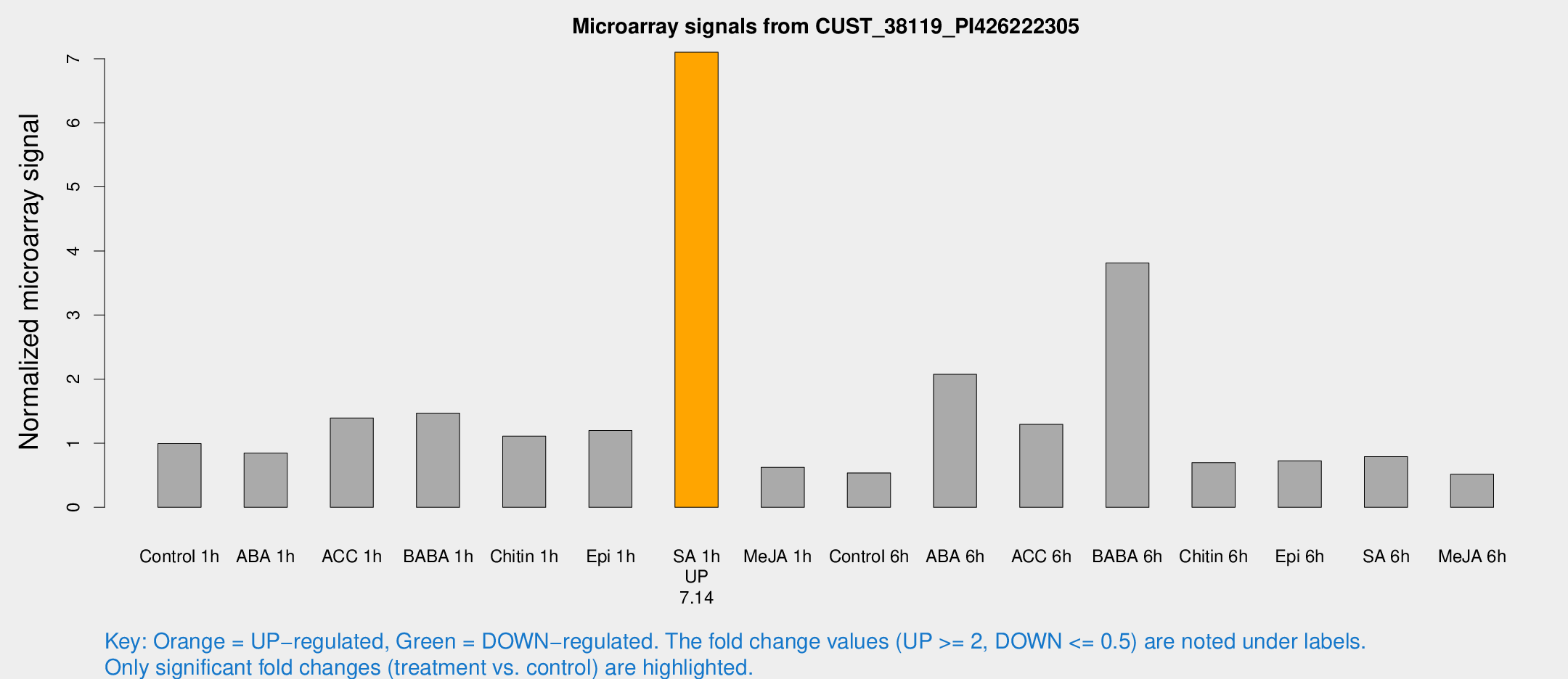
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 254.019 | 45.0123 | 0.993923 | 0.106428 |
| ABA 1h | 186.113 | 11.1932 | 0.847032 | 0.0808015 |
| ACC 1h | 391.132 | 106.351 | 1.39285 | 0.369958 |
| BABA 1h | 358.187 | 50.8378 | 1.46994 | 0.144897 |
| Chitin 1h | 249.627 | 23.302 | 1.10902 | 0.0655779 |
| Epi 1h | 257.366 | 15.2084 | 1.19688 | 0.0716004 |
| SA 1h | 1995.37 | 547.214 | 7.10056 | 2.16452 |
| Me-JA 1h | 128.658 | 17.9065 | 0.622259 | 0.0410751 |
| Control 6h | 156.067 | 55.6442 | 0.536041 | 0.189388 |
| ABA 6h | 599.111 | 171.843 | 2.07404 | 0.555772 |
| ACC 6h | 371.254 | 31.8258 | 1.29525 | 0.294276 |
| BABA 6h | 1096.67 | 213.676 | 3.81175 | 0.646661 |
| Chitin 6h | 184.594 | 11.8486 | 0.696188 | 0.0548951 |
| Epi 6h | 224.037 | 73.4139 | 0.726112 | 0.226424 |
| SA 6h | 195.771 | 17.7774 | 0.790982 | 0.0479557 |
| Me-JA 6h | 140.468 | 45.9164 | 0.516562 | 0.127081 |
Source Transcript PGSC0003DMT400054793 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G50930.1 | +1 | 2e-139 | 419 | 235/496 (47%) | cytochrome BC1 synthesis | chr3:18929817-18931547 FORWARD LENGTH=576 |
