Probe CUST_37796_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37796_PI426222305 | JHI_St_60k_v1 | DMT400077055 | CCTACACGGCATATTATCTGAAATATGTCAGGTAAGATGTGTACATCTACTTGTTCAATT |
All Microarray Probes Designed to Gene DMG400029967
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37776_PI426222305 | JHI_St_60k_v1 | DMT400077054 | CCTACACGGCATATTATCTGAAATATGTCAGGTAAGATGTGTACATCTACTTGTTCAATT |
| CUST_37796_PI426222305 | JHI_St_60k_v1 | DMT400077055 | CCTACACGGCATATTATCTGAAATATGTCAGGTAAGATGTGTACATCTACTTGTTCAATT |
Microarray Signals from CUST_37796_PI426222305
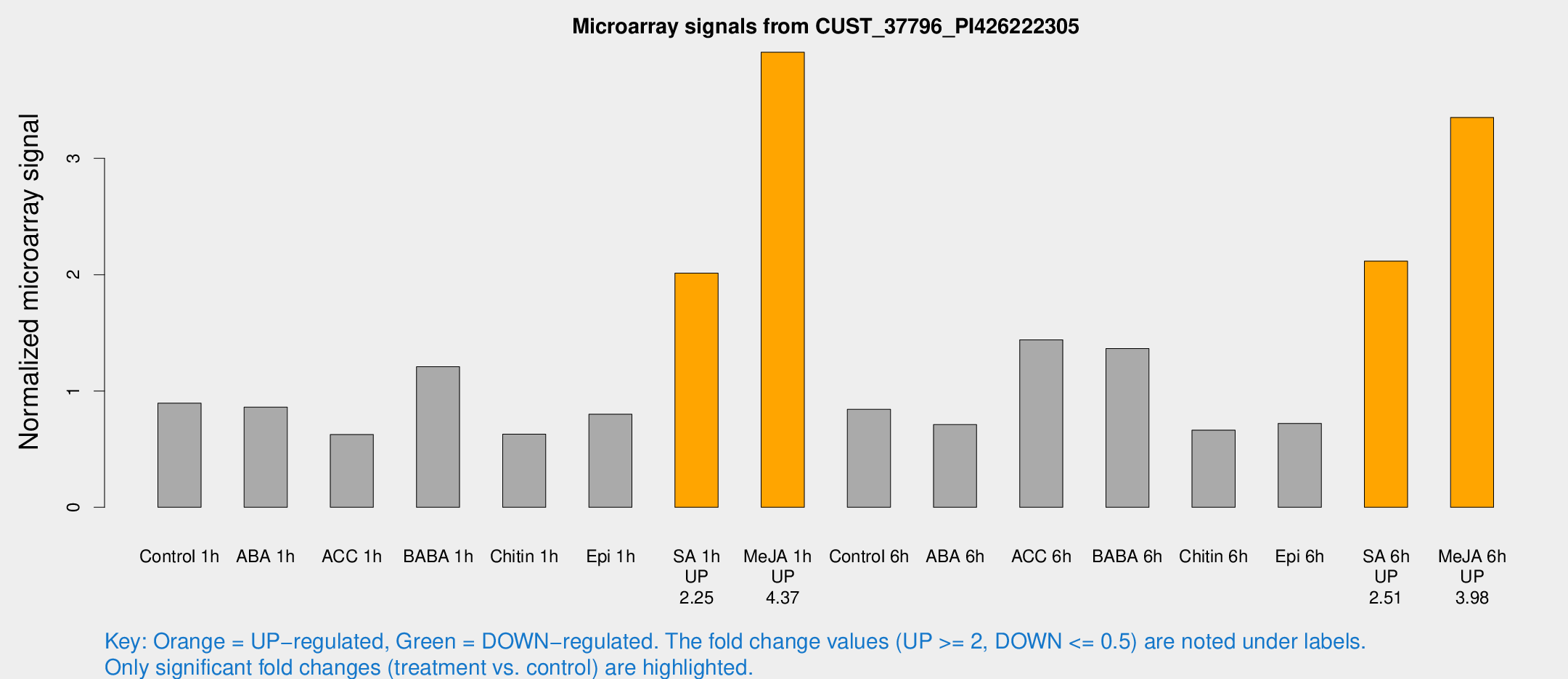
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 837.445 | 209.998 | 0.895755 | 0.159655 |
| ABA 1h | 824.829 | 324.515 | 0.861221 | 0.415503 |
| ACC 1h | 748.272 | 389.009 | 0.625367 | 0.355539 |
| BABA 1h | 1096.11 | 250.373 | 1.20828 | 0.202092 |
| Chitin 1h | 520.177 | 112.722 | 0.628898 | 0.0891098 |
| Epi 1h | 617.565 | 65.635 | 0.799734 | 0.0805767 |
| SA 1h | 1876.98 | 308.876 | 2.01271 | 0.284942 |
| Me-JA 1h | 3012.29 | 816.532 | 3.91238 | 0.693182 |
| Control 6h | 813.278 | 225.85 | 0.841763 | 0.189141 |
| ABA 6h | 695.24 | 141.205 | 0.711175 | 0.110055 |
| ACC 6h | 1503.04 | 268.797 | 1.43992 | 0.370978 |
| BABA 6h | 1398.51 | 267.932 | 1.36441 | 0.268392 |
| Chitin 6h | 622.607 | 36.1138 | 0.663378 | 0.0384622 |
| Epi 6h | 732.678 | 108.672 | 0.72123 | 0.0888635 |
| SA 6h | 1971 | 449.658 | 2.11647 | 0.325646 |
| Me-JA 6h | 2948.53 | 170.671 | 3.35167 | 0.264972 |
Source Transcript PGSC0003DMT400077055 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G10600.1 | +1 | 2e-139 | 421 | 221/517 (43%) | cytochrome P450, family 81, subfamily K, polypeptide 2 | chr5:3351227-3352777 FORWARD LENGTH=516 |
